In a Rabbit Hole
But not in a cave
Greetings substack audience.
I have been taking a bit of break from X and Substack to explore a new method for DNA sequencing. I have not retreated from this fight. I was in fact in DC last week to speak at the MAHA Institute and had the pleasure of meeting Wafik El-Deiry, Del Bigtree, Nicolas Hulscher, MPH, and Maria Gutschi in person. It was great to finally meet many I only know online.
However, I was not born to tear things down and live in a perpetual state of protest. I need to build. A perpetual screaming into the void destroys your ingenuity, erodes your soul, while hopelessness becomes a contagious nihilism. Additionally, I have found the medical freedom movement to enter the “eat each other” stage of divide and conquer and its becoming an unproductive time sink for me.
el gato malo nailed it in this post.
One afternoon, someone asked me a question about lagging strand synthesis and this made me realize most of the patents I was awarded for SOLiD sequencing are now open source and after 20 years there are so many better ways to improve upon this method with new enzymes, better detection methods, and pretty much every proprietary reagent we relied on back then, now over the counter and available by multiple competitive vendors.
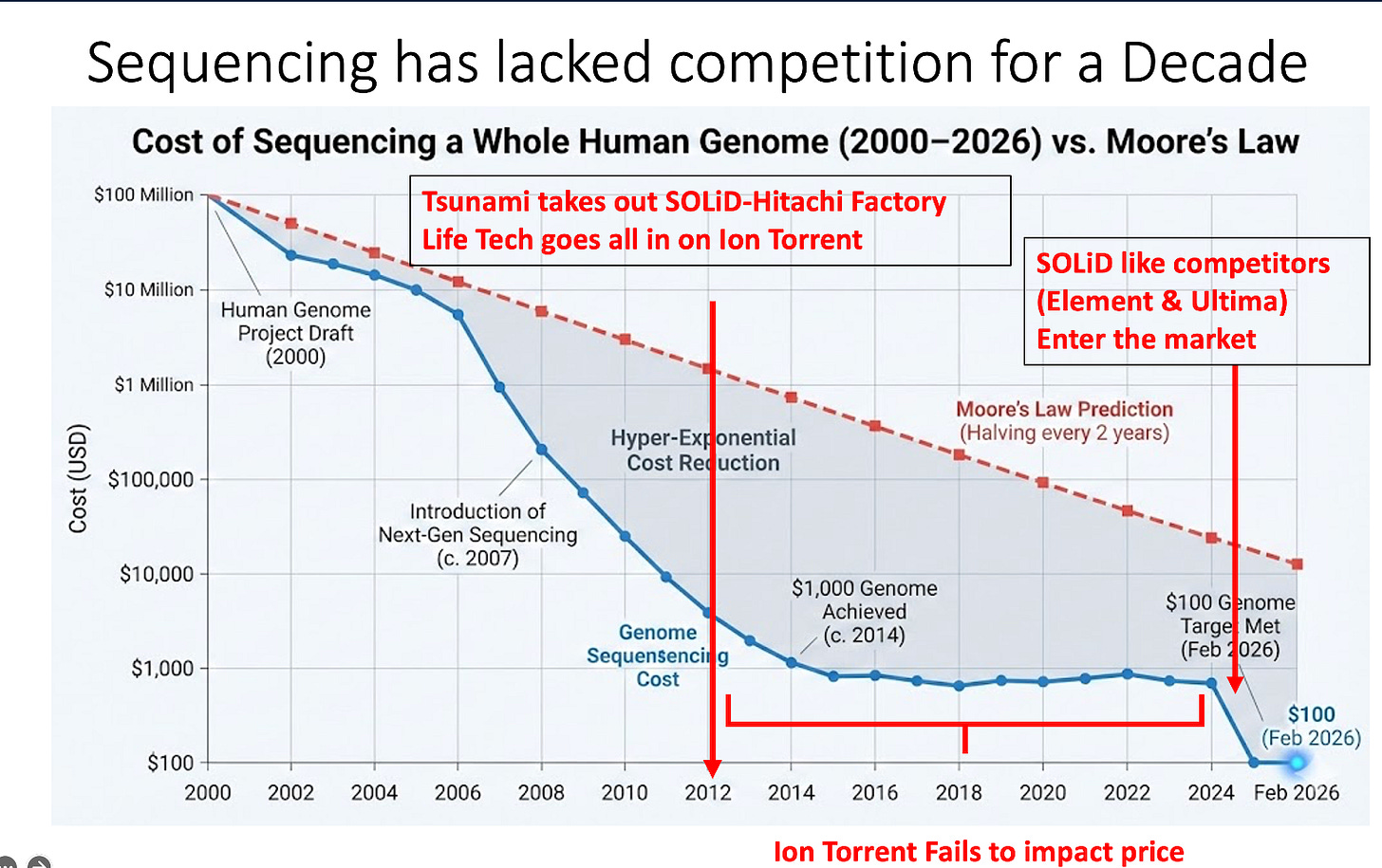
As I watched the 2026 AGBT presentations roll out I was struck at how the price declines in DNA sequencing came to screeching halt after the SOLiD Hitachi plant was taken out by the Tsunami in Japan. At this point Life Tech was forced to go all in on Ion Torrent with hopes of it giving Illumina a run for its money. It fit a niche and had its own remarkable achievements with brilliant people pushing it to the extremes but for a decade Illumina held a near monopoly and price declines didn’t materialize.
Now some of Illumina patents are also expiring and this has opened the door for 2 promising SOLiD/ILMN like competitors to enter the market. Element Bio and Ultima Genomics have come to market with their own cheaper versions of Sequencing by Synthesis and prices are beginning to fall again. Ultima Genomics looks like FISSEQ on beads deposited on circular wafers. Think sequencing on a 30 tray CD player.
Element did something very clever and instead of relying on a single fluorophore per base, created avidity tags that allow them to decorate 5-10 fluorophores per base and improve signal to noise.
While Oxford Nanopore and PacBio have made groundbreaking advancements on Long reads, 80% of the market doesn’t need these as they are working with fragmented DNA in FFPE or liquid biopsies. For most human genomics markets, high read count matters more than the long reads that dominate the smaller de novo sequencing market. The market seems to have settled on 2X150 base pair reads being sufficient for most applications and while several companies offer more expensive 2x300bp reads, they are rarely run as often the DNA is shorter than this or the improvements don’t justify the longer run times or sequencing costs.
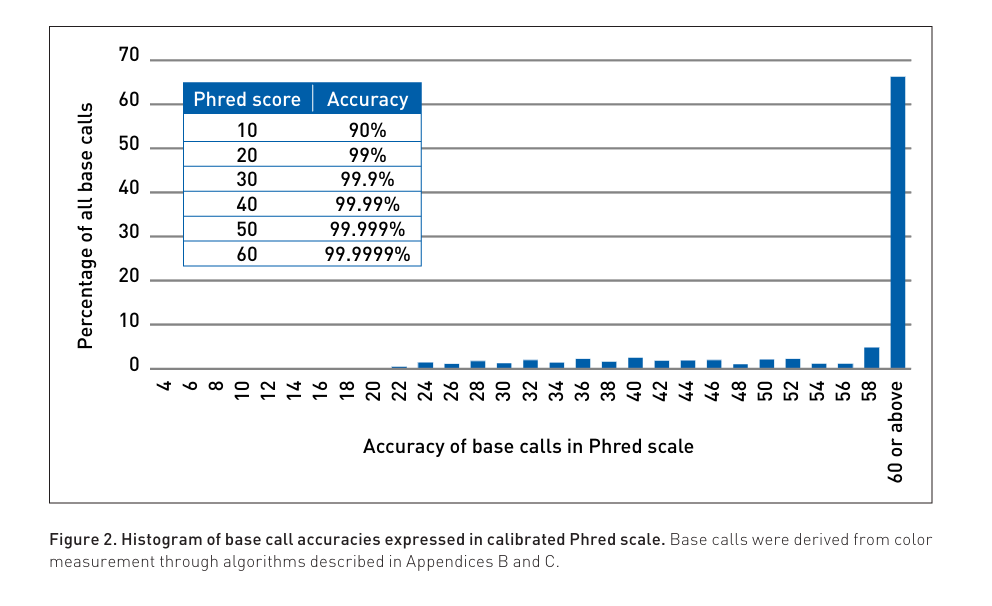
Now the entire competitive race appears to have shifted back to accuracy. Q30 reads to Q40 reads are now the race.
As of this date no sequencer has matched the accuracy of SOLiD. This was largely due to its use of ligases that don’t incorporate a single base but 5 bases at time so very interesting error correction codes could be engineered into the dye read out strategy. SOLiD just suffered from Short reads (75bp)…. 20 Years ago.
If you are into Galois fields and Claude Shannon like error correction in DNA sequencing this is a good place to start but this unique system saw the the first next gen sequencing data that was capable of sensing 1 in a million polymerase errors. A 2x150 sequencer with SOLiD like accuracy would be the ideal combination today.
The new sequencing method we are exploring certainly shares many of these themes with SOLiD and may in fact turn out to be an excellent approach for Next Gen sequencers we’ve brought to market before but that is not our immediate focus.
There is a gap in the sequencing market that often leaves people overly trusting qPCR data. In many ways this plagued the COVID era as the Drosten PCR approach failed to perform this level of validation before sparking a public panic with false positive C19 PCR assays that had no internal controls and many errors in their primer sequences. I’m not a virus denier and I use qPCR everyday but qPCR can easily lead to over confidence in conclusions if you don’t sequence validate what you amplify and now we find ourselves in a world were a few shoddy qPCR asssays can lead to global lockdowns, culling of millions of animals and vaccines heavily contaminated with DNA as regulators never scrutinize the veracity of the qPCR results being produced by their sponsors.
For this reason, many people, including the inventor of PCR , Kary Mullis, prefer qPCR data be sequence validated before screaming “Pandemic”.
But this step often gets skipped as NextGen sequencers cost over $1000 to run. They certainly deliver an enormous amount of data for this $1000 but it is CostCo like batch economics. You achieve low cost by buying in bulk and dropping $1000 on every $10 PCR amplicon makes no sense. You need to barcode these samples and pool them with others PCR Products to multiplex them on these large volume sequencers to make any economic sense. This batching process ultimately delays turn around time as you wait for enough sample pools to make these expensive runs make economic sense.
Sanger sequencing often fills this niche. It’s a $billion a year market, still growing 3%/year but there are only ~50,000 Sanger instruments globally. They are $50-$150K per instrument and require weekly watering to prevent the Capillaries from drying out. As result they tend to be centralized in facilities and require shipment to these facilities to run single amplicons. Usually a 48 hour Turn Around Time and $10-$20/read after shipping is included.
Oxford Nanopore (ONT) looked like a nice solution to this application space with their $90 Flongles but even these need 10 amplicons barcoded to get the costs below $20/sample. Libraries need to be made and pooled to make the chip costs get close to Sanger sequencing. The library construction and barcoding costs bring in another ~$20 per sample and this becomes cost prohibitive unless you need deep sequencing data for each amplicon. Most dont need this level read depth to verify an amplicon.
Very recently they replaced the CEO of ONT and now the lower throughput/lower cost P2S platforms are being sunsetted due to the high support costs of the lower throughput markets.
Our solution to this market gap is a method than can run on the 500,000 -1M qPCR instruments already in the field post COVID and only require a small liquid handling robot to automate the methods.
The patent application has been filed but the exact details I will reserve for a later post when we have more data in hand. At the moment, we have several specialty reagents being synthesized to get this proof of concept nailed down.
We can see this being helpful in many markets where a sample is PCR positive but people want sequence confirmation of the amplicon to speciate it in an afternoon. Given the chemistry behaves on single amplicons, it is entirely congruent with running in massively parallel systems we have designed before only with longer readlengths and some of the error correction capabilities built in.
The NextGen market isn’t our immediate focus as we want something to address our current customer base using qPCR in the Cannabis, Food and Supplement testing space. The capacity to sequence a PCR amplicon in an afternoon without any additional equipment other than the qPCR instrument already in place and a liquid handler most labs already have, would certainly change our lab and very likely many others.
Stay tuned. As this develops, I’ll be certain to keep this audience apprised of the Journey.





It's a bit hard to follow being that I'm an amateur but so fascinating.
Thank you.
"I have not retreated from this fight."
THANK YOU!
Bwsides, I never thought I'd ever write a "ME TOO!"