The critics of our work have focused a lot of their negative attention on the average fragment size of the DNA in the vaccines.
Lets just remind the reader that now the EMA, the FDA and Health Canada have acknowledged that SV40 promoters are in fact in the vaccine. Despite some of them accusing us of lab contamination, they now can see the sequence in the data Pfizer gave them. They only decided to look 3 years later after a few billion doses. This is materially different than them actually picking up a pipette to confirm this. They simply opened a dusty file and realized.. holy shit, McKernan and Buckhaults are right, and in some cases like Health Canada, they admitted this was an underhanded move by Pfizer.
Handing someone a long list of ATCGs is a snow job. You need to annotate what the sequence means and in this case Pfizer chose to selectively annotate the plasmid, effectively deleting the presence of the SV40 components that would be by default annotated until 2 years later when some undergraduate pot head researcher decided to sequence it.
Yes, Factcheck.org had to remind their audience that I have an undergraduate degree in Biology while mentioning my response to their questions was thrown on Twitter. They of course didn’t link to my response on twitter as their fact check ultimately failed to address my response.
A follower of mine posted a great fact check from June on this topic and compared it to the same fact check from October. Its like they can’t even fact check their own fact checks fast enough to keep up with how many lies they must weave to support the narrative. It appears many people have caught onto this.
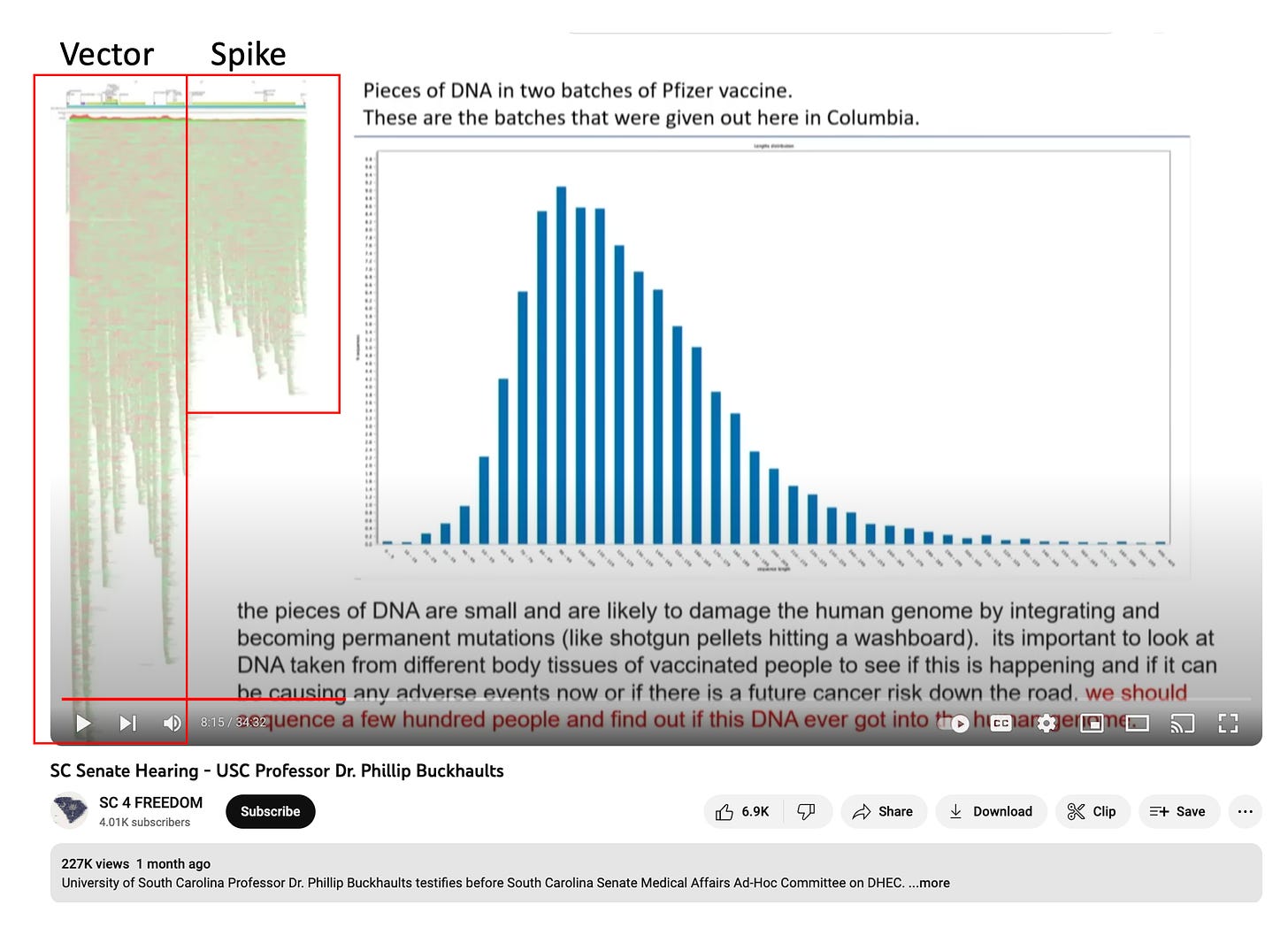
So why does the fragment length of the DNA matter and why are the nudge nannies zeroing in on this? There are FDA regs that point to 10ng/dose that is >200bp. Other people from the FDA (Klinman et al) are concerned about smaller fragments (7bp) being meaningful if they are biologically active sequences like promoters. The vax-optimists like to claim that these small fragments will all be destroyed in the cytoplasm and are meaningless, but they are making a common error in statistics related to measuring averages.
Mean fragment lengths are deceiving in non normal distributions. Any kurtosis or Fat Tail effect really distorts these numbers. We are going to touch on the impact of this in measuring DNA fragment sizes but first we need to assure you that artifacts that have influenced the DNA coverage maps of these plasmids in the past are now addressed.
In the past, leaving the RNA in the vial while making Oxford Nanopore (ONT) DNA sequencing libraries gave us less sequence coverage over the Spike regions, presumably due to the RNA interfering with the DNA ligation of the spike DNA. See our sstack on DNA:RNA hybrids.
This DNA:RNA hybrid effect can be seen in Philip Buckhaults sequencing coverage which used a sequencing method that left the RNA in place (No RNase). The sequence coverage over the vector (No RNA) is 2X as deep as the coverage over the Spike region (lots of ModRNA). If we dont address this skew in coverage, we will likely have a biased estimate of the DNA fragment sizes.
So to get rid of the RNA…
I took all the vaccine available (~1ml) in high adverse event Pfizer lot FL0007. Treated this with RNase and 55C for 120 minutes to eliminate the RNA. I also ran a time course of this RNA decay and it degrades in 1 minute at RT. I was bit surprised at how quickly this destroyed the RNA but the RNase stock I used was 20mg/ml. So 50ug-100ug of RNaseA was added to 10ul of vaccine and decayed this in under a minute.
This is really important as it shows how much modRNA inflates the DNA estimates using Fluorometry (AccuGreen) quantitation of the DNA. Removing the RNA with RNaseA knocks the DNA signal down a log scale but we are still a LOG scale over the 10ng/dose limit. We mentioned this potential intercalating dye crosstalk in our preprint with
. Now we have it measured and it doesn’t get Pharma out of the woods.After this RNase treatment, there should be no RNA inhibiting the ligation reaction or influencing the sequencing coverage.
Oxford Nanopore sequencing of high adverse event Pfizer lot FL0007.
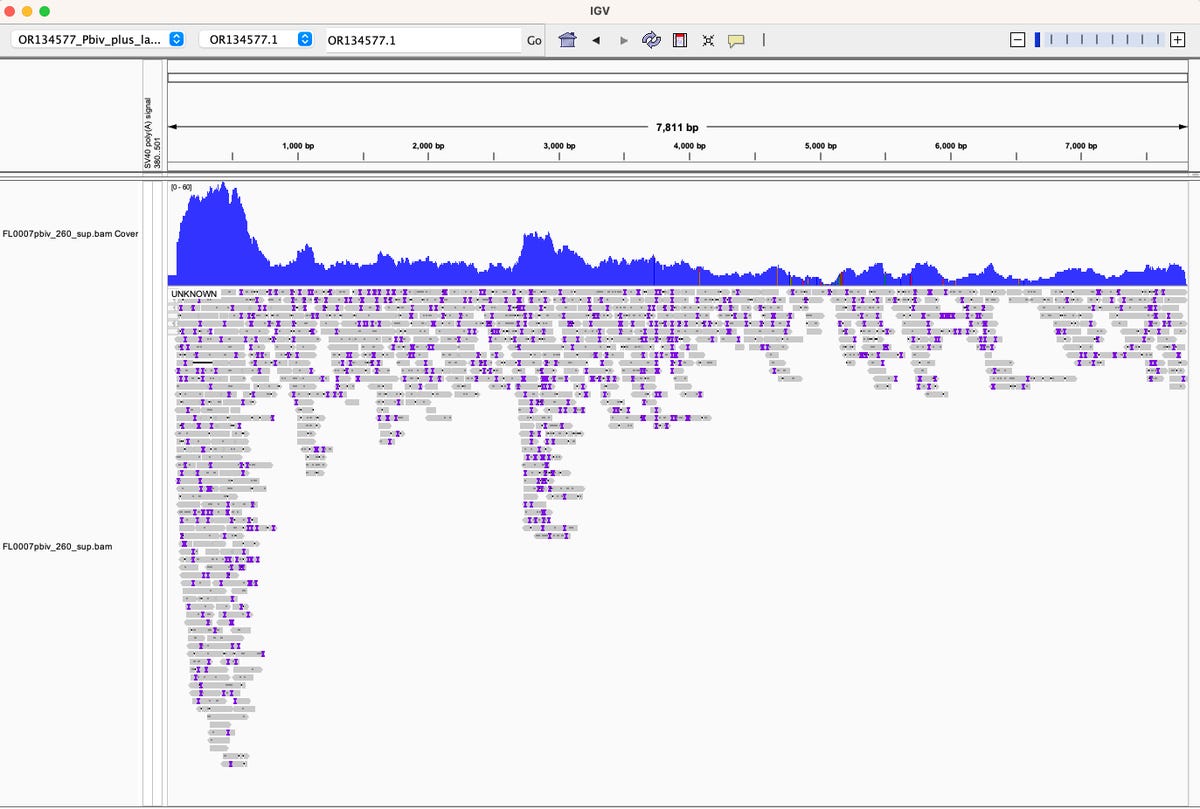
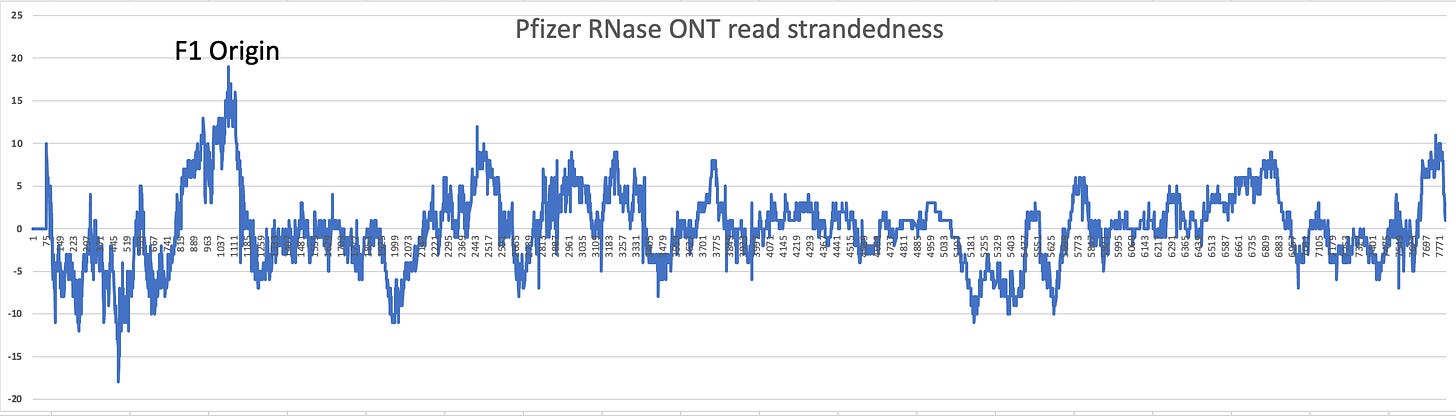
The high coverage on the left side of the coverage map is the F1 Origin.
The read strandedness (Watson:Crick ratio) is skewed in this location. In normal DNA libraries your Watson:Crick read ratio should = 1. When an origin of replication is actively making ssDNA, this read ratio departs from 1. This implies the F1 origin may be active in their manufacturing process. This needs to be assessed with an orthogonal technique to confirm.
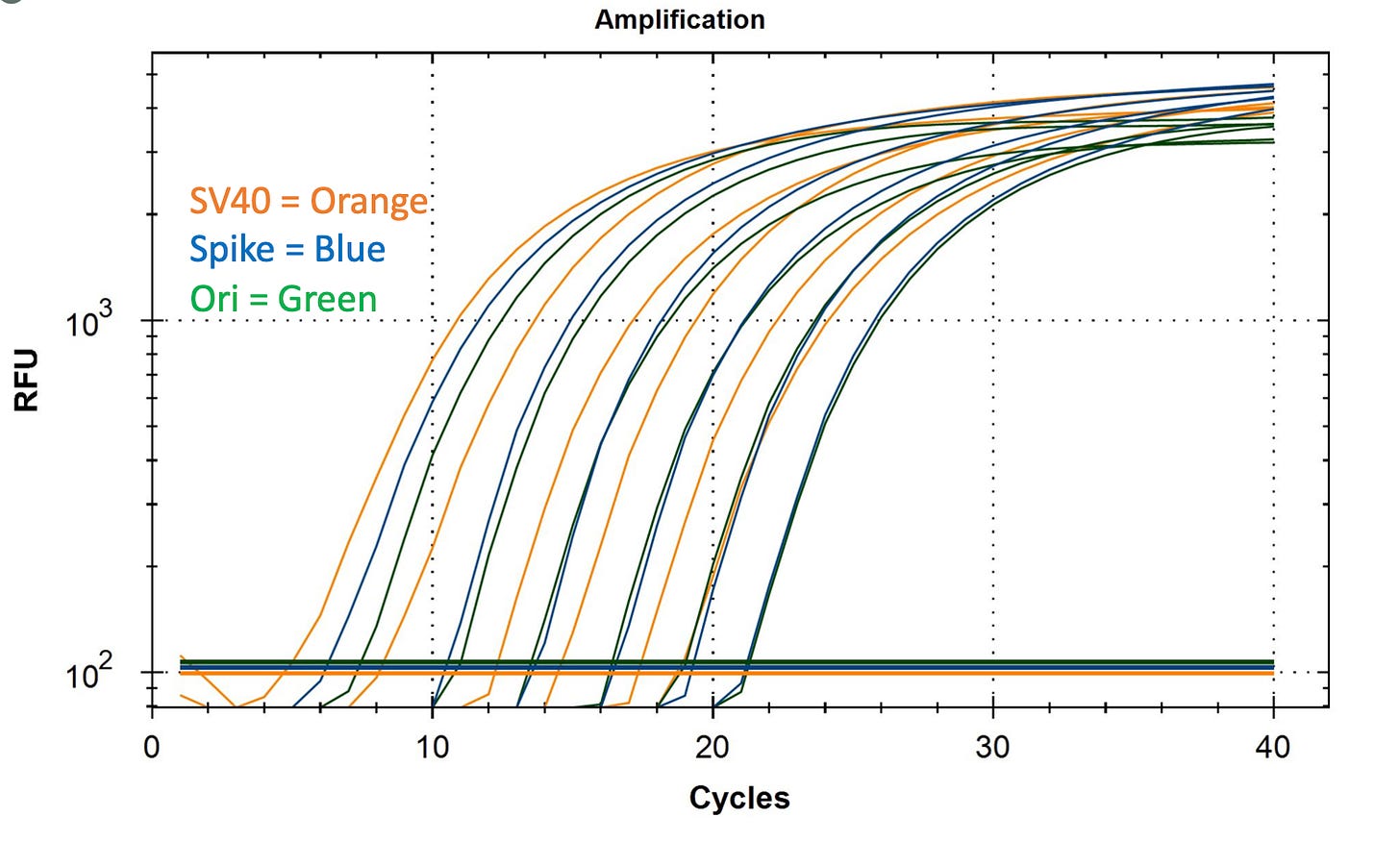
The qPCR DNA levels of these vaccines clocked in at CT 17 in a triplex assay (all assays combined and competing).
Standard Curve for SV40 uniplexed compared to Spike and Ori Duplex (Not in a Triplex reaction). Some performance changes occur when assays are triplexed. SV40 DNA is from IDT gBlock.
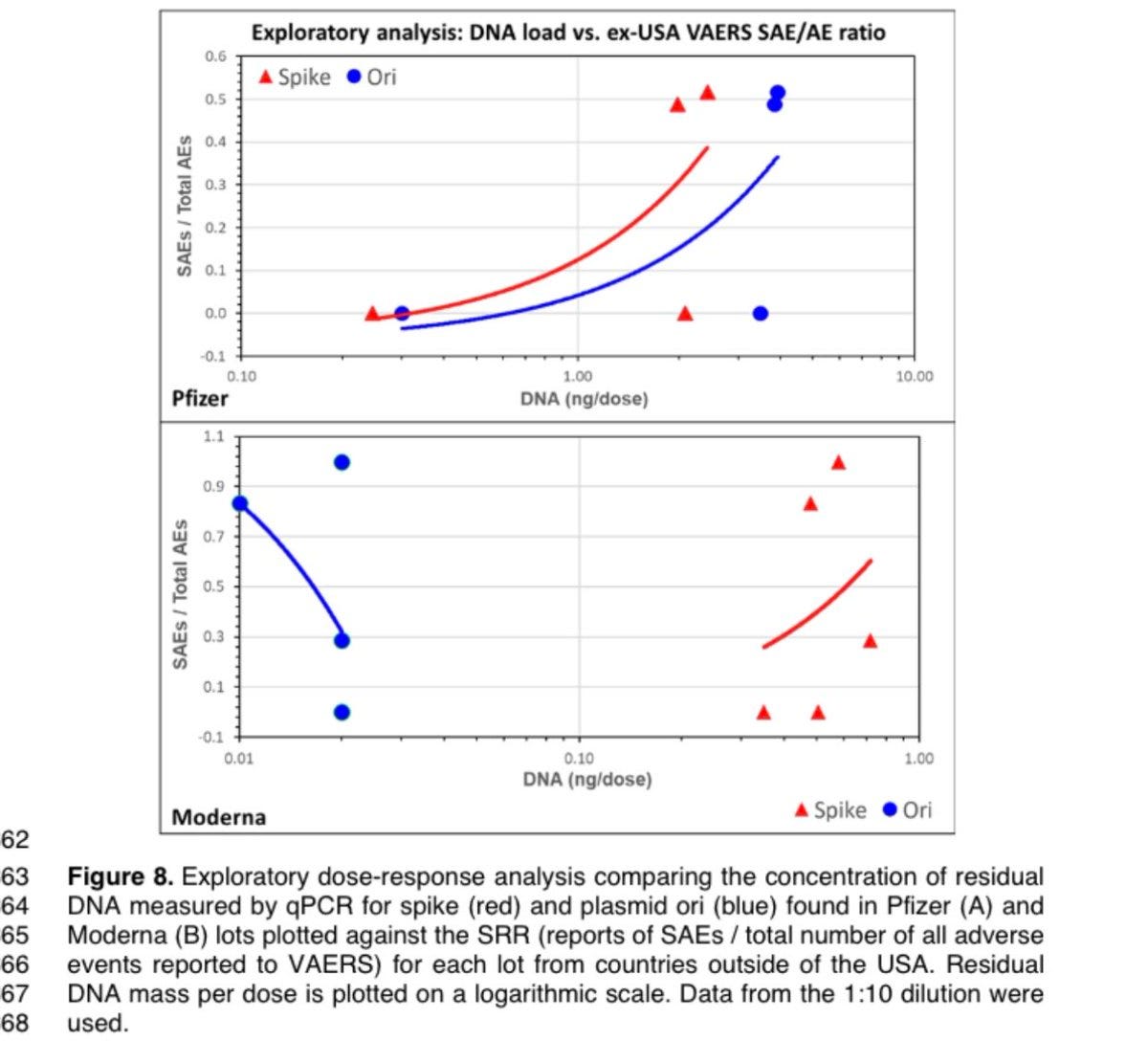
FL0007 is high in AE and high in DNA. They would be on the upper right part of the Pfizer graph by @AdhesionsOrg
Howbadismybatch
So we have a high concentration lot with 100s of Billions of DNA fragments per dose, supported by both qPCR and RNaseA treated DNA fluorometry. The lot has High Adverse events. Does this shrapnel mean anything?
Back to Kurtosis-
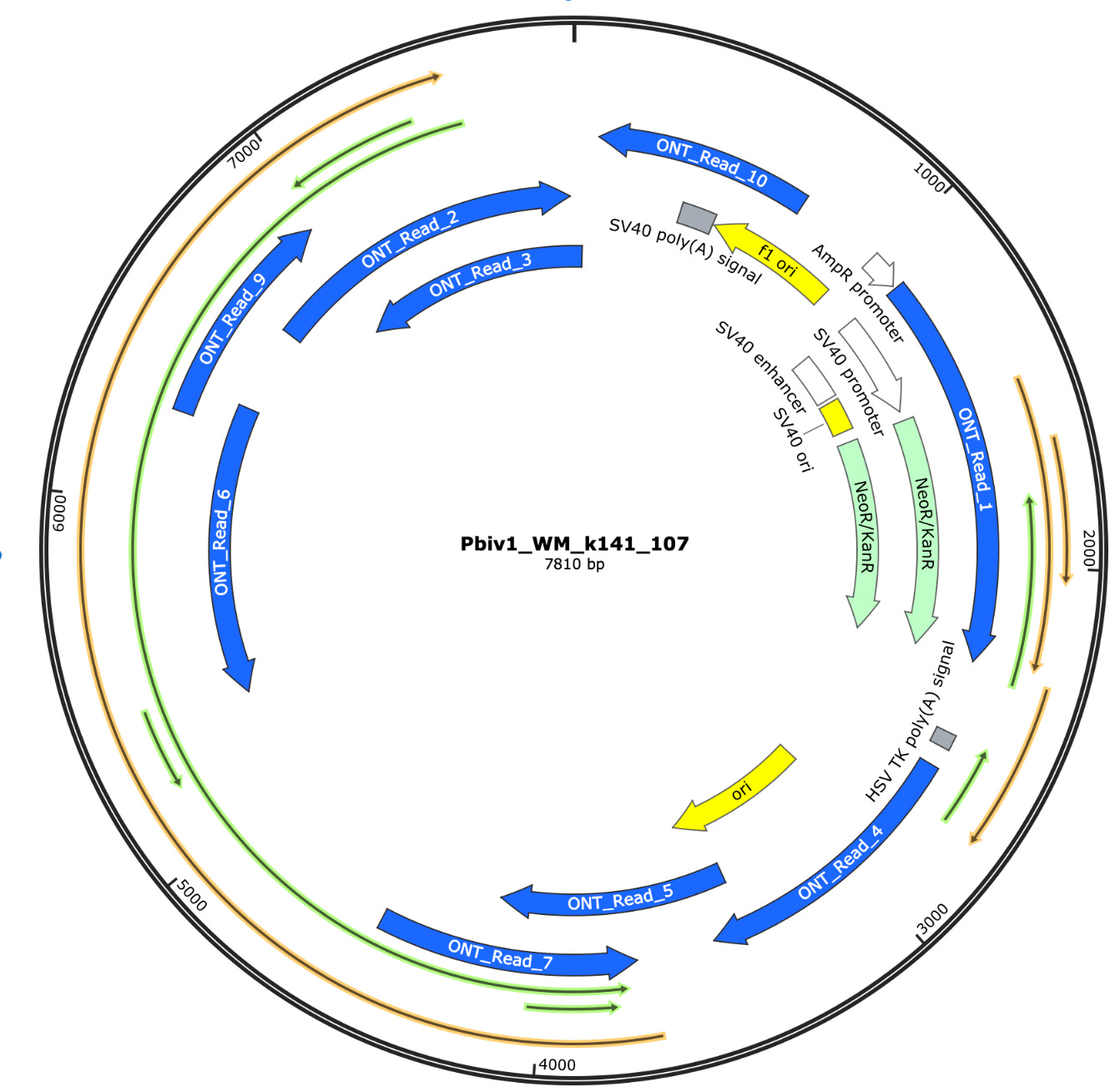
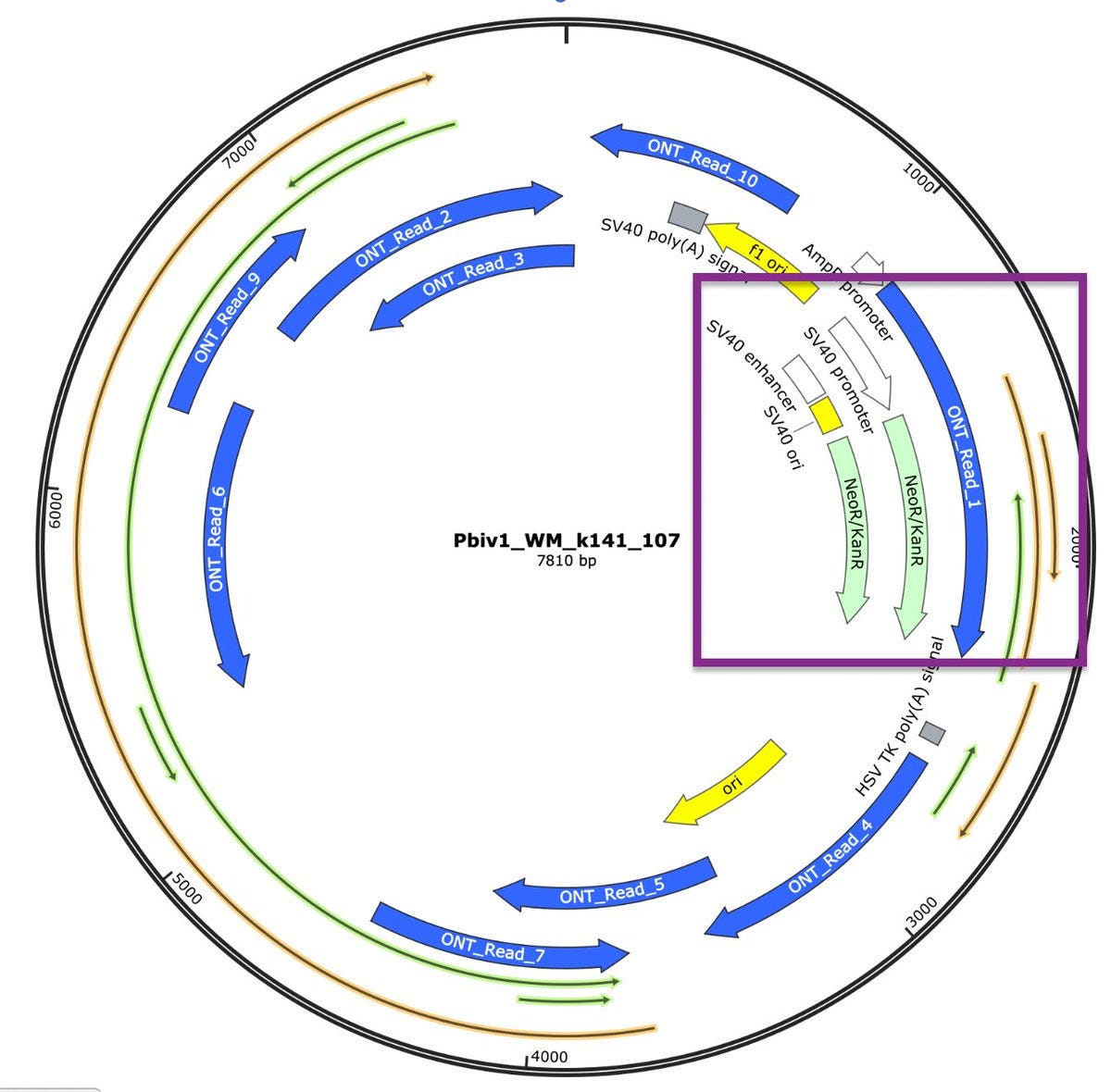
The 10 longest reads from a 612 read ONT Flongle run are depicted in blue on the map below. 1 in 60 reads are over 630bp, and there are 100B molecules per dose... That’s 1.6 Billion molecules per dose over 630bp. Does that sound like its broken up too small to matter? One read in Purple spans the entire SV40 promoter and Neo/Kan gene.
There is another detail in here. The EMA is now claiming that SV40 promoters were left out as they are immaterial to plasmid manufacturing.
Doah…That’s not only not true but it is an embarrassing disrobing of the emperor.
You need a promoter of the Neo/Kan gene or no plasmids get made! The exact opposite of what they have claimed is in fact true. Yes, it’s true the SV40 promoter is redundant to the AmpR promoter seen in Pfizer but the Pfizer plasmid map given to the EMA omitted the AmpR promoter as well. How can they claim the promoters are immaterial to plasmid manufacturing if there is no selectable marker expressed? No selection = No plasmid. It gets lost in E.coli’s rapid cell division.
AND IT GETS WORSE!
This won’t get them out the woods as Pfizer also failed to identify the 2nd largest ORF (Open Reading Frame) in the plasmid. This is the mystery ORF of 1252 amino acids described by Beaudoin et al.
ORFs are painted in Green and Gold in the above plasmid map. What is the Mystery ORF in Green on the opposite strand of Spike? Why was this ORF never disclosed despite the WHO and FDA regulation claiming all ORFs need to be identified? This seems like a major omission as FDA can’t claim this is of no consequence if regulations demand all ORFs be disclosed.
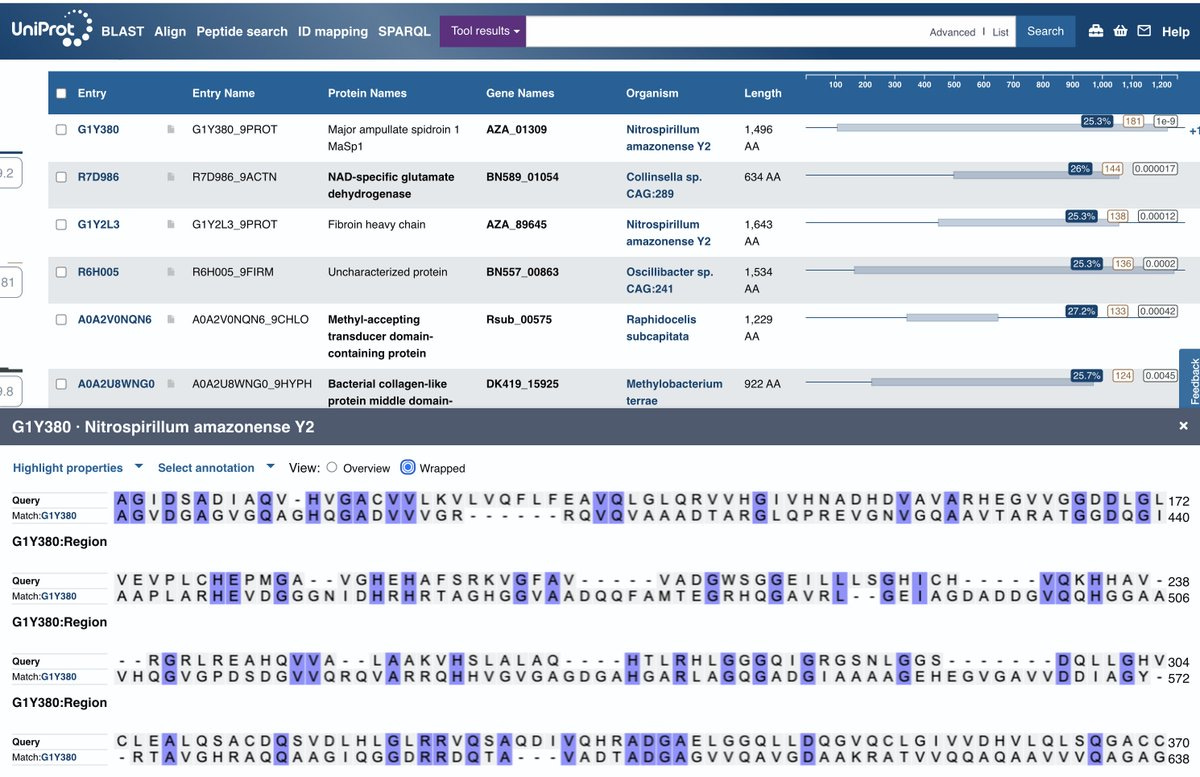
That Mystery ORF's closest hits is to a protein involved in silk.
Other hits with lower E-values are to Collagen and Fibroin. I get no hits when I blastP this sequence against NCBI. It only shows up in UniProt and the E-values (E-9) are nothing like what you get from blasting the Spike strand but its the only thing that shows up.
Silk gene paper link attached.
https://www.nature.com/articles/s41467-022-32093-7
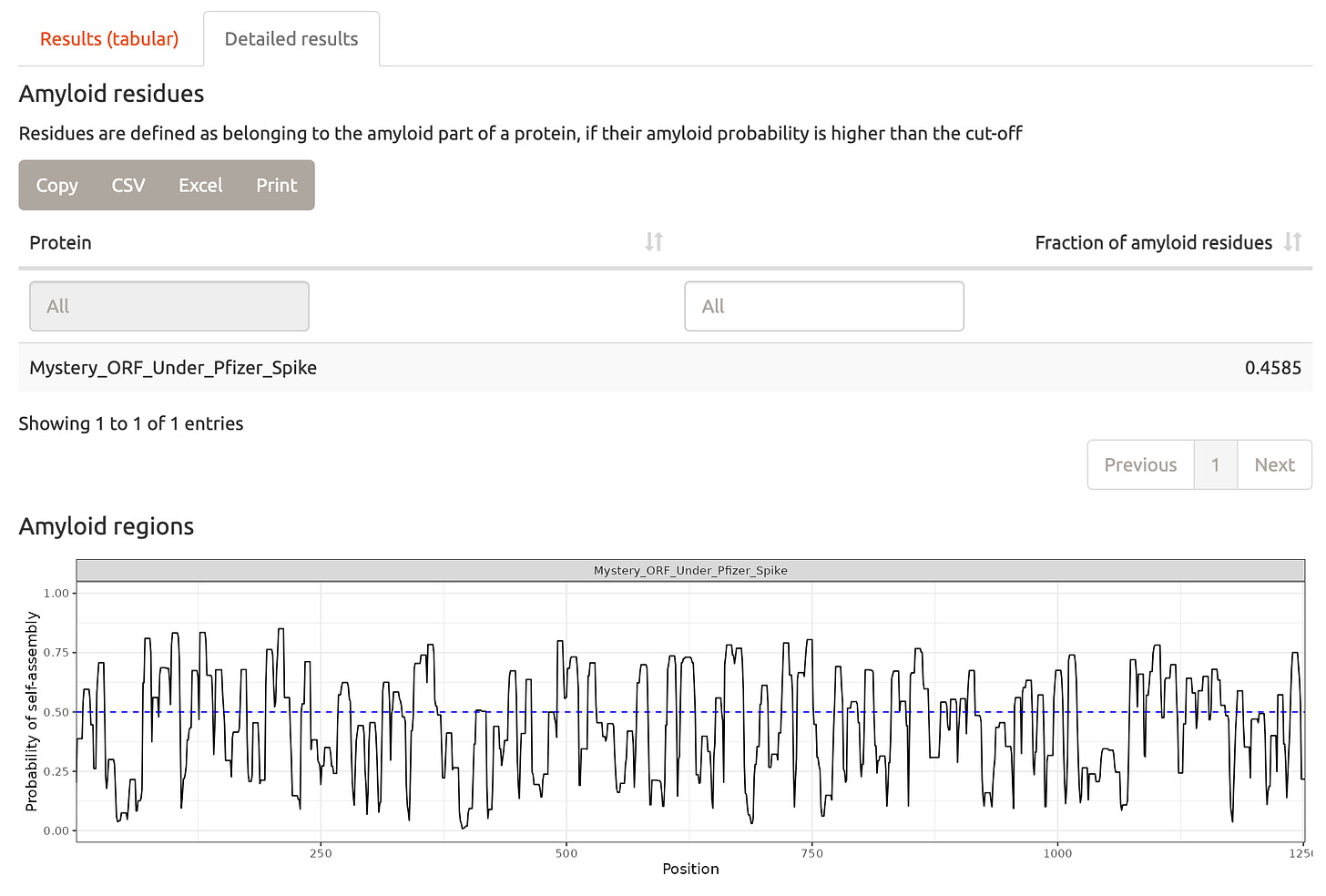
These silk proteins have many amyloidogenic domains.
WHO and FDA documents declaring all ORFs need to be identified.
The EMA is clearly confused over the presence of the SV40 promoter/Ori/Enhancer. Their language regarding its role in plasmid manufacturing is self contradictory. It appears they haven’t loaded this sequence into SnapGene and should. Once they do this, they will realize there is a large 1252 amino acid ORF that is undisclosed and this violates FDA and WHO guidelines. This ORF may be amyloidogenic.
has done some preliminary work to show that amyloidosis is more frequently reported in Pfizer than Moderna.Does this mystery ORF express?
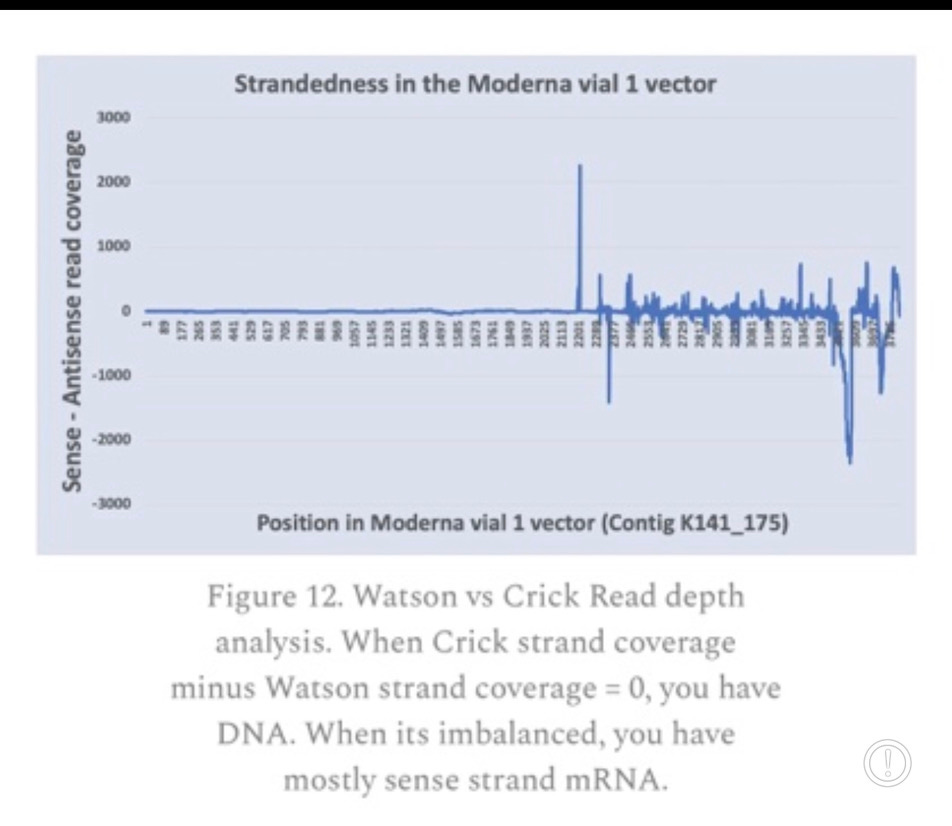
The SV40 Promoter/Enhancer is a bidirectional promoter so its possible RNA is transcribed. In support of this, we have previously reported RNA-Seq data of these vaccines that demonstrated very high levels of Watson strand RNA in the vaccine (blue line that drops down below the Zero line in the figure below). It should all be Crick strand (Sense) RNA but the RNA-Seq doesn’t support this. The T7 promoter is the tall peak at 2200 on the chart below. Everything to the right of that peak should all be Crick strand RNA and above zero. Several regions are negative and are Watson strand (anti-sense) read mappings implying dsRNA which is coding on both strands with the Mystery ORF. Keep in mind this strandedness analysis is on a Moderna bivalent vaccine and so the coordinate system isn’t the same as the Pfizer map above.
The other possibility causing the transcription of this mystery ORF is that T7 Polymerase strand flips when making the 1273 amino ORF onto the other Mystery ORF strand. During the process of making the RNA from the DNA, the polymerase decides to switch templates. Like a photocopier getting confused on printing on both sides or one side of a page, these polymerases can sometimes meet a RNA molecule that hairpins on itself due to palindromes in the sequence and flip strands and start making RNA in the reverse direction. In this case the reverse direction encodes an amylodogenic peptide.
Codon optimization Ooops.
Many people scream that strand flipping is so rare and that this hypothesis is a stretch.
It is so much of a stretch that Moderna engineered a T7 polymerase that does less of it…
They chose to get this done in 2023. Maybe they learned something.
I guess we will have to comb through Mass Spec data to see if any of these reverse strand mystery ORFs turn into proteins.
We shouldn’t have to do this. We have a failure to disclose by Pfizer of an ORF involved in silk, collagen and fibroin. Mysterious clots are getting pulled from patients arteries.
When will the state regulators flex their 10th amendment and ignore the dereliction of duty from the bought and paid for federal regulators? Hopefully somebody in these federal agencies is a good soul and will rattle the cage from within but the signaling we see from these agencies, is that they will be deemed ‘Safe and Effective’ regardless of any body count you show them. We are only beginning to understand the mechanisms but the forensics on their approval process reveals too much deception for them to defend.
This was recently covered on The Defender.
The protein sequence of the Mystery ORF is
>Mystery_ORF_Under_Pfizer_Spike
MQYQLESSCVVQFHALQHGLRIVLVELAAAATATTALQAATAAGHATQHDCDHHDGNQSGDKAQPDVPGPLDVLLVLPQFLQVDQALVQILGHLVQPVDLFLDVHDAGIDSADIAQVHVGACVVLKVLVQFLFEAVQLGLQRVVHGIVHNADHDVAVARHEGVVGGDDLGLVEVPLCHEPMGAVGHEHAFSRKVGFAVVADGWSGGEILLLSGHICHVQKHHAVRGRLREAHQVVALAAKVHSLALAQHTLRHLGGGQIGRGSNLGGSDQLLGHVCLEALQSACDQSVDLHLGLRRVQSAQDIVQHRADGAELGGQLLDQGVQCLGIVVDHVLQLSQGACCAAQAVLDLADGAVELVGDQLLVLVQHILGHSDAVEPVGHLHSKGDLQSGACSKCPAACDCAGQQGRCVLGDHLIGQQRRQHCQSVKLLGANQIPGGNVAQTIAILLDEAGVGQCHFVEQQVLDEAPLAGLARIGQNLAEIEAAEVLDRRGLVDLLHLGEHLLGVLVLFHGDPCQGSFQLGAEAAVLQQQVGALGGIAADVHGAVHAGLGHGHRQDLCGHADGEVGGDSDRVVGVGHAVLGAQRHCVGNDALAGHASGSPVALCLCLVAGADSSADGDVALVAIVHVLGSDQTAGSGLKHIAAGGVHPPCRCQLIGVNGHGHFGTVHALVQHCHLIAGVGARGDHRHSAEAARGDVQDFQCLGISNGVCGIGDIPAKLLEWQELLVALCQHAGAGQAVEVEVHAFVLHEIGAFLRAAHCGRGMQQFEAQHHHSVGLVAHAVCGPKAVGLQWEVAVHACHAVTRLVAGLIDLGGDVPLEGLQIGLPEQPVPVIVVAADFGVQLVAVPGNHTAGEVVGQLVVVVGDVACLSRGNLPHFVSPDHEAVGVHVCEAQVVQLGRGHAVALECEEGGEVVQHGVVGHAIADPLPVPGVHRGESGGIEHLVEGAQIGDIGEPHDGFGGLHPEVAGLVDALFHGEGLQGALCLAQRIQSTIHGVGDGAVLVVLQQEGSRLQVAHIVSGGTSCPSAAAIARCQVASVQGQQCLKPGDVDADGQIHQGFQSREALRQIPAEVDRGVLAVDLEVAVDVLKHELAQVLEVALLAFQVHQERLGHVLEGAVVGAAVHPELAFHPALVVLVVVDVQEGVVAELELAHFDDHVGGVVHDQQALGLAVQCGAEDPASDDVGLLGAGKVHPVVEGQHGVVESLGAIGAGDGVEPGHVAEERQEQVLGRVQHAGSEHLVGVVHASGKAVGV*
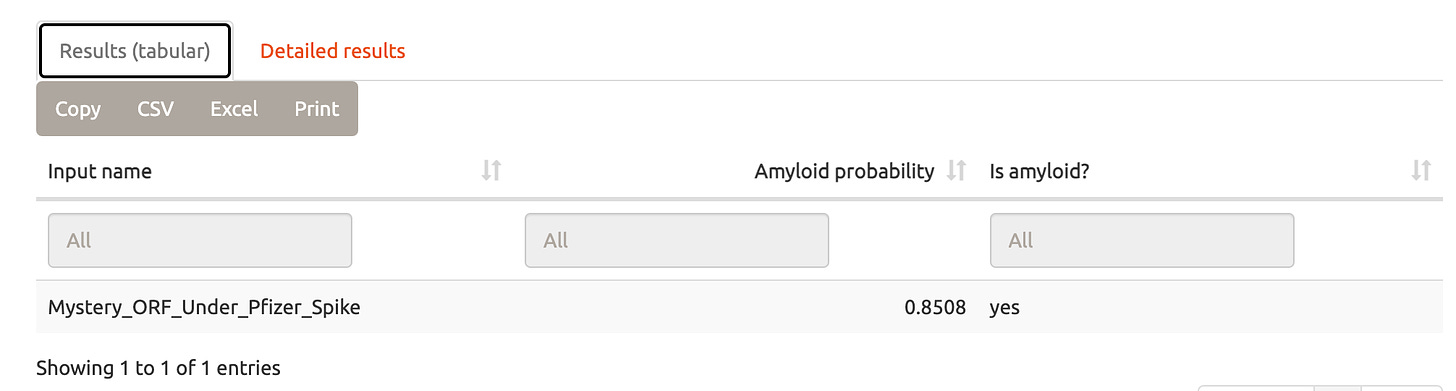
Amyloid prediction-
http://biongram.biotech.uni.wroc.pl/AmyloGram/
Mystery ORF
Pfizer Spike























When evidence is presented from multiple parties that there are lots of problems, like death, with these injections and the regulators do nothing for years how is this explained? The obvious answer is the people running the regulators and thus the senior management at the regulators are both involved in the fraud. It will take time for enough people to accept this depressing reality. We all need to believe our own eyes, and face up to the ugly truth. Some will never face reality, there are a lot of cowards, but there are enough people that want the truth.
"..we shouldn't have to do this...." true. Glad you are though!