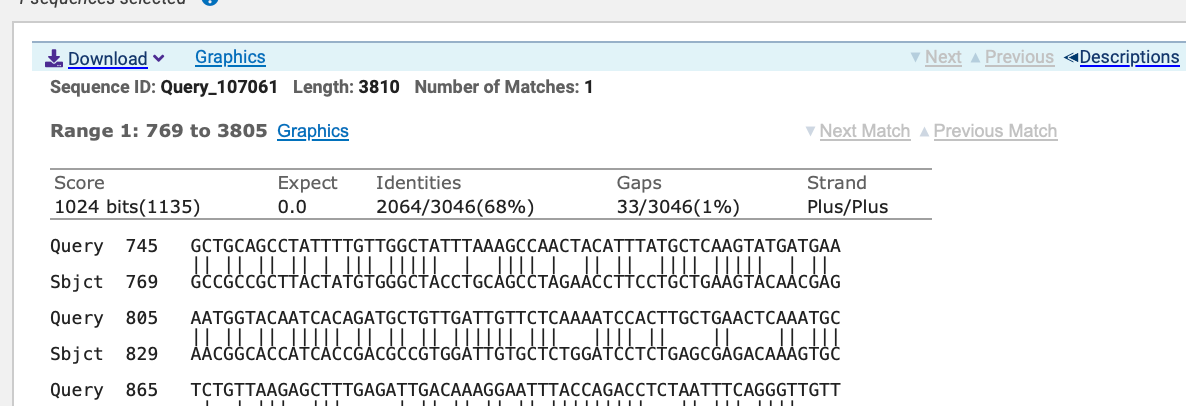
A curious follower on twitter took to BLAST (Credit to Mabel_Syrup_) after seeing the Spidroin work. This wasn’t their background but they were a quick learner. They noticed when you took the Spirdroin ORF we published and BLASTed it using DELTA-BLAST against the patent database you can find some hits to a SARs1-Tor patent. US patent 7267942 filed in March of 2004.
It takes some time to dig through the patent office and find the 2470 amino acid sequences they stuffed into the patent so I have pulled them down and linked them here (credit to Mabel_Syrup_ for sleuthing these).
What on earth are they doing with 2470 genes in a corona virus? This doesn’t make any sense.
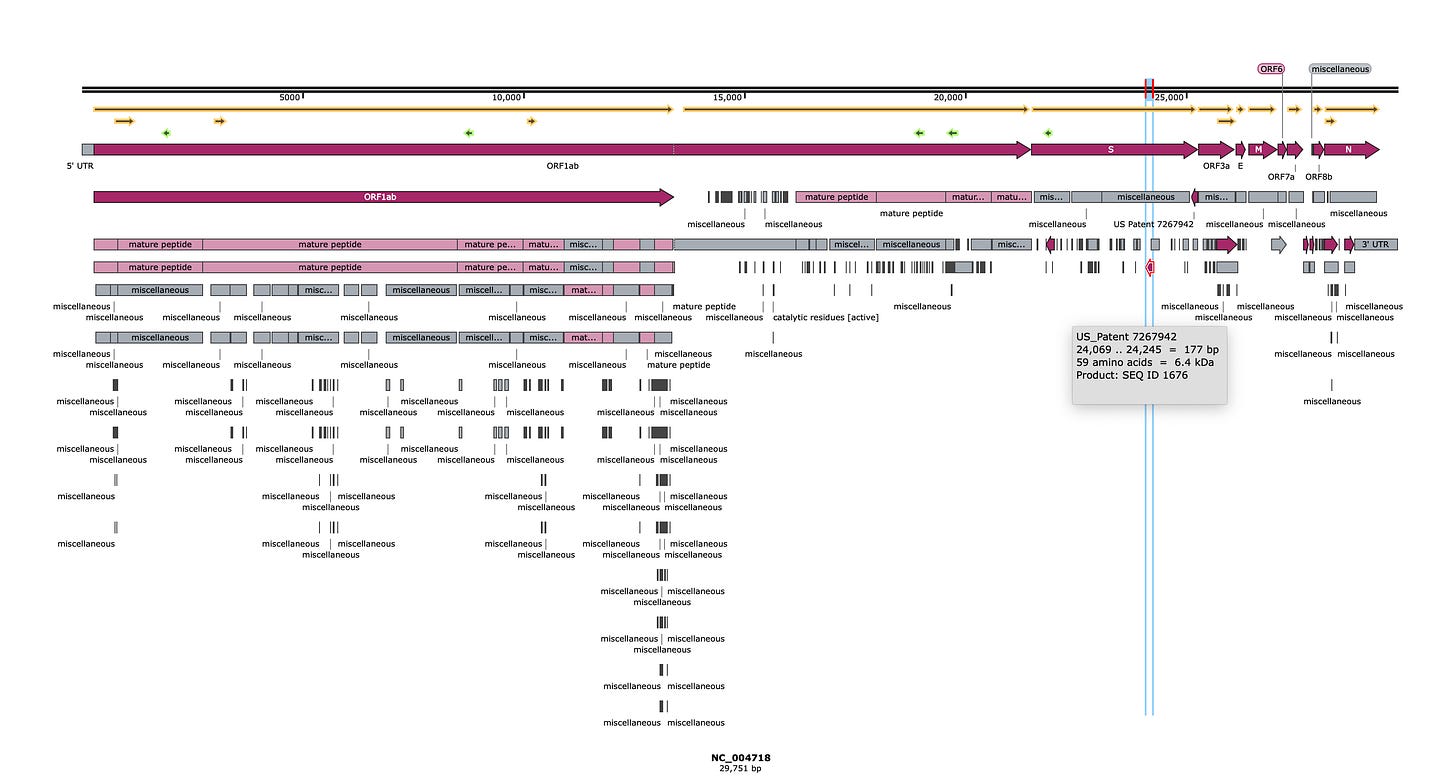
So I download the genome (NC_004718) into SnapGene to have a look.
It appears this is the first sequence of SARs1 and the patent attorneys in their rush to file IP, didn’t know where the genes were so they did what any sloppy and greedy patent attorney would do.
They 6 way translated the 29Kb sequence and filed on every ORF they could find with in-silica prediction tools.
It was so sloppy they even filed on 4 amino acid ORFs!
It was so sloppy (or greedy) they even filed on ORFs on the reverse strand that would never exist in a single stranded RNA virus. They would exist in a dsDNA plasmid attempting to encode that virus but thats a different story.
Not all of these sequences are claimed in the claims of the patent. They exist in the specifications of the patent so if the inventors ever find use for them, they can go back and file continuation in parts or divisional applications and have the benefit of the earliest priority date.
The Pfizer vaccine should have some similarity to SARs spike as it is a codon optimized SARs2 spike.
So we should expect SARs1 peptides to match Pfizer Spike and its reverse complement ORF to some degree.
The sequences in this patent that have some homology to Spidroin are sequences 1659-1701.
Lets walk through them and visualize them in SnapGene so you can see what is going on. You will need the NC_004718 reference listed above.
As you search SnapGene for each of these micro-ORFs you can see they are all in frame on Spidroin but there are Stop codons interrupting these micro-ORFs from becoming a full 1252 amino acid Spidroin ORF.
Some how Pfizers codon optimization ablated all of these stop codons?
How did that happen?
Well first you have to understand that this spidroin ORF is not only on the opposite strand but its perfectly in frame. There is no Frame shifting.
What does that mean?
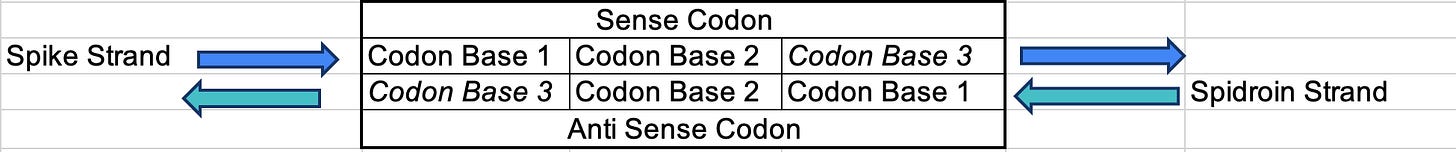
Each Spike strand codon has 3 bases.
The 1st base on the spike strand = the 3rd base in the Spidroin codon. The 3rd base in each codon is the most degenerate base. Its called the wobble base for this reason. There is most freedom to change the 3rd base without changing the amino acid so codon optimization tools are mostly changing Base 3. This happens to be base 1 in the codon for Spidroin.
Since Pfizer was busy changing their 3rd base in most codons, and they avoiding using any Leucine or Serine codons that could form a Stop codon on the opposite strand… Voila! The eliminated ~40 stop codons from SARs1-Tor to make a 1252 amino acid open reading frame under spike.
If you back up and look at the whole Spidroin ORF in SARs1 you can see its fragmented with many stop codons. Between each yellow and green ORF is one or more Stop codons.
At least 39 Stop codons where ablated by Pfizer to make a full length 1252 amino acid ORF.
Keep in mind the virus is single stranded RNA so it does not have this template despite the patent attorneys including these fragmented microORFs in its patent specification.
What is bizarre is how Pfizer managed to resurrect this into a full length 1252 amino acid ORF and failed to disclose it to regulators who demand all such open reading frames be disclosed in their submission.
Here is the SnapGene file for anyone who wants to take a look.







Wait. Is it possible that Pfizer reused someone else's patented DNA?
You say "It takes some time to dig through the patent office and find the 2470 amino acid sequences they stuffed into the patent" but then in the next para you say "What on earth are they doing with 2470 genes in a corona virus?"
Did you mix up genes and amino acids?