Intent to Deceive
A 20 minute tour through Snapgene
This is a tutorial on how to use an open source plasmid annotation tool known as SnapGene. Dr. Byram W. Bridle has mentioned he also uses this software tool to work with his plasmids. Its quite common because it is great software and they get you hooked with a free trial.
We are going to use the Pfizer plasmid sequence as tutorial.
We are going to ask SnapGene to open the sequence file that was likely given to regulators.
You are going to see how SnapGene by default will annotate the plasmid with SV40 components and the work you would need to do to remove it.
1)Download a free trial of SnapGene. There are some functions that are limited until you pay for it but I think the annotation tools will be live. Install it and open it. If the free version doesn’t offer this feature, let me know and I will film this process for those who don’t have access.
2)Download this Fasta file of the Pfizer plasmid sequence. Remember where you download the file as you will need to find it to direct the Open Files….menu in SnapGene.
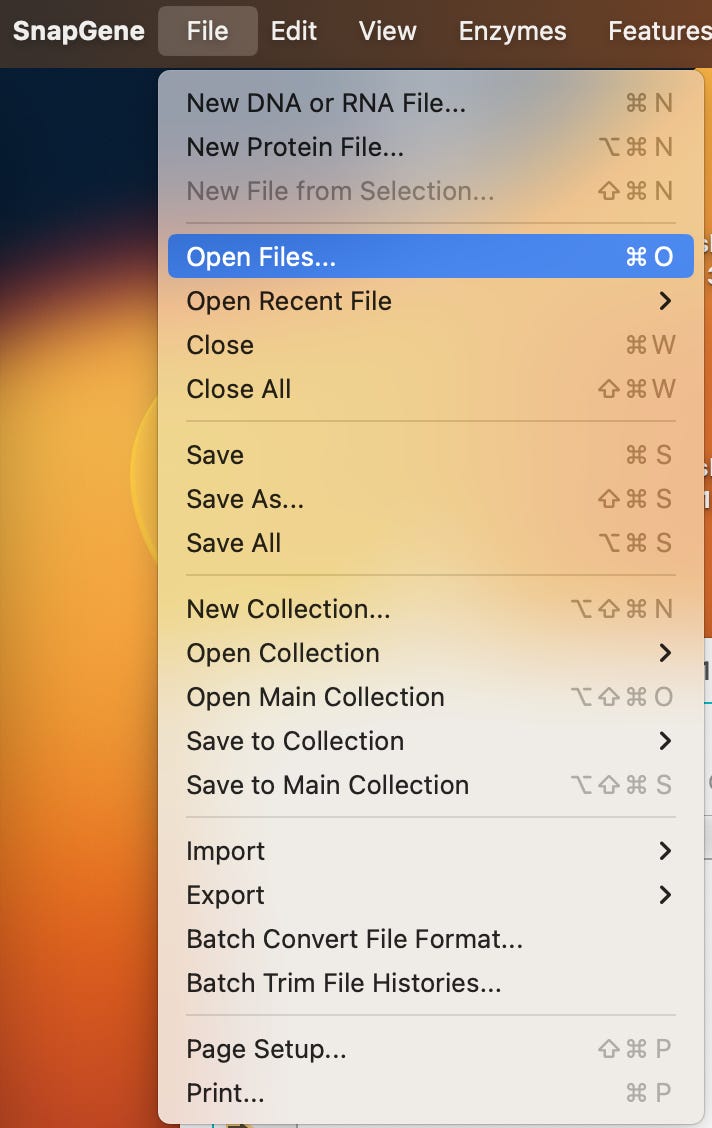
3)Select on the menu Open Files…
Select your file.. it should be named Pbiv1_WM_k141_107.fa in the folder where you saved it.
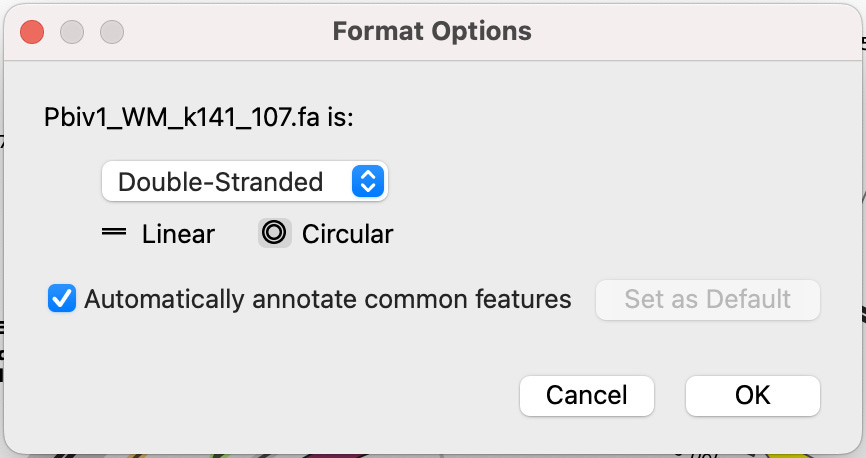
It will ask you a few questions on Formats. Select Double-Stranded and Circular. Check the “Automatically annotate common features” check box.
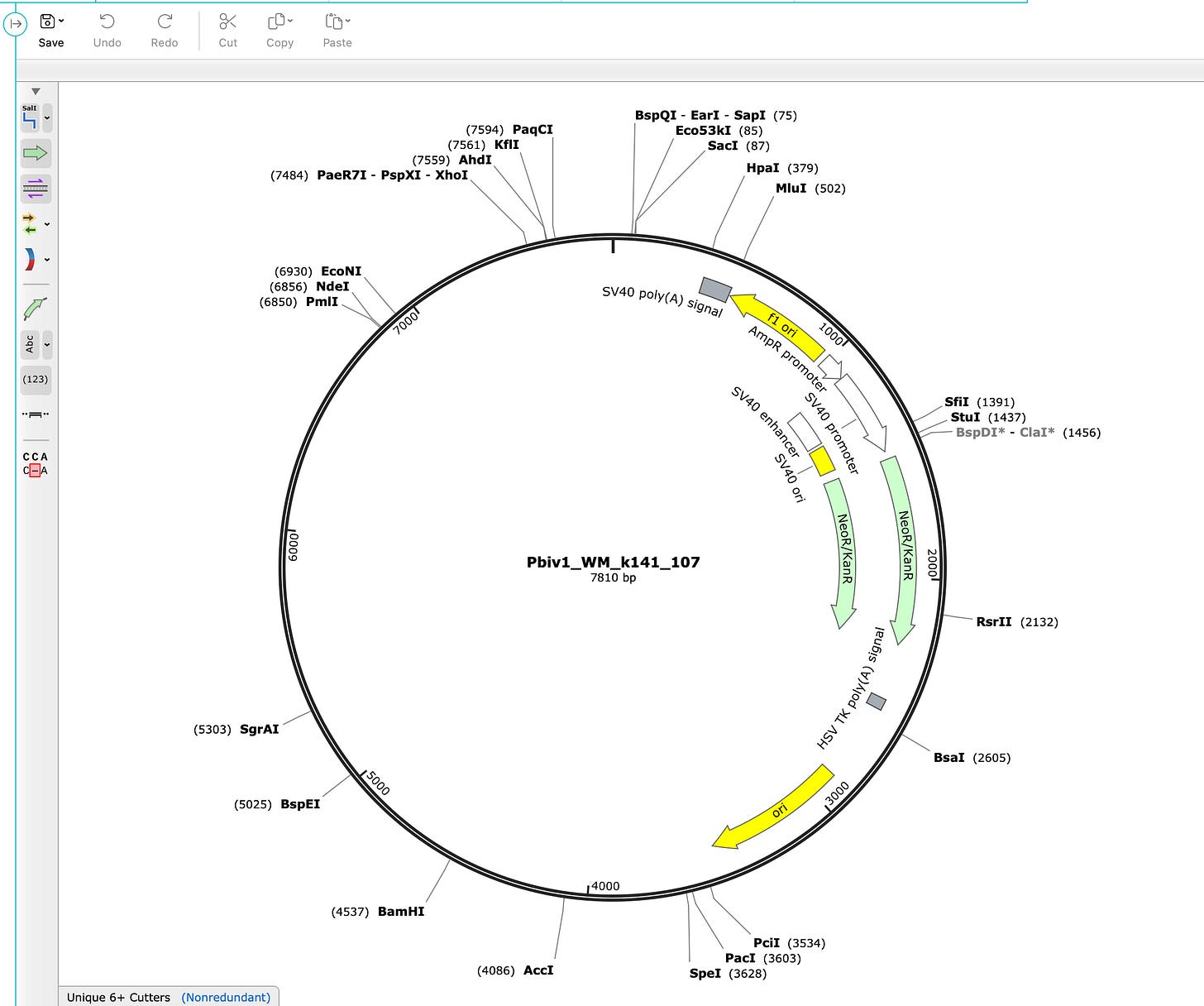
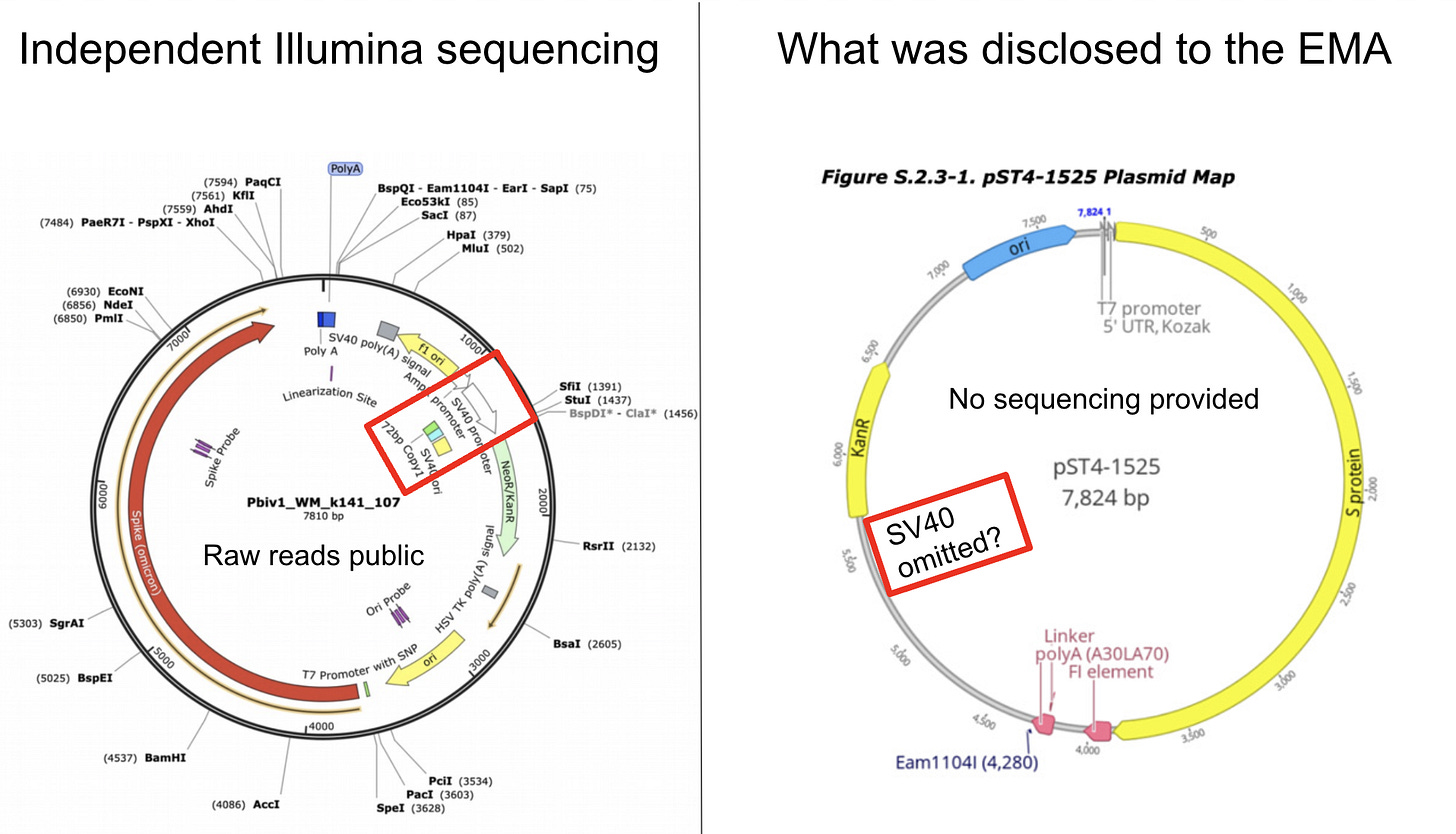
A plasmid map will appear like the one below.
Note how this industry standard tool annotates from 5 o’clock to 1 o’clock the Bacterial Ori, The Neo/Kan gene, The SV40 Ori, The SV40 Enhancer and the SV40 Promoter, the AmpR promoter, The F1 ori and the SV40 polyA Signal with zero effort.
It does not annotate the spike region as it has no way of knowing what this is until you teach it.
Now to get to the plasmid map Pfizer gave to the Regulators you would have to surgically delete the SV40 components. Note Pfizer had to annotate the plasmid with details as small as the 5bp Eam1104I site and the T7 Promoter and the Kozak site… all features smaller than the 366bp SV40 Promoter/Enhancer/Ori. Any tool that ran this auto-annotation would have also labeled the SV40 region.
So someone had to actively delete this region before handing this to regulators. This is no accident. Someone spent time to manually delete something.
This is intention to deceive. This is legally different than “Ooops, we forgot that”.
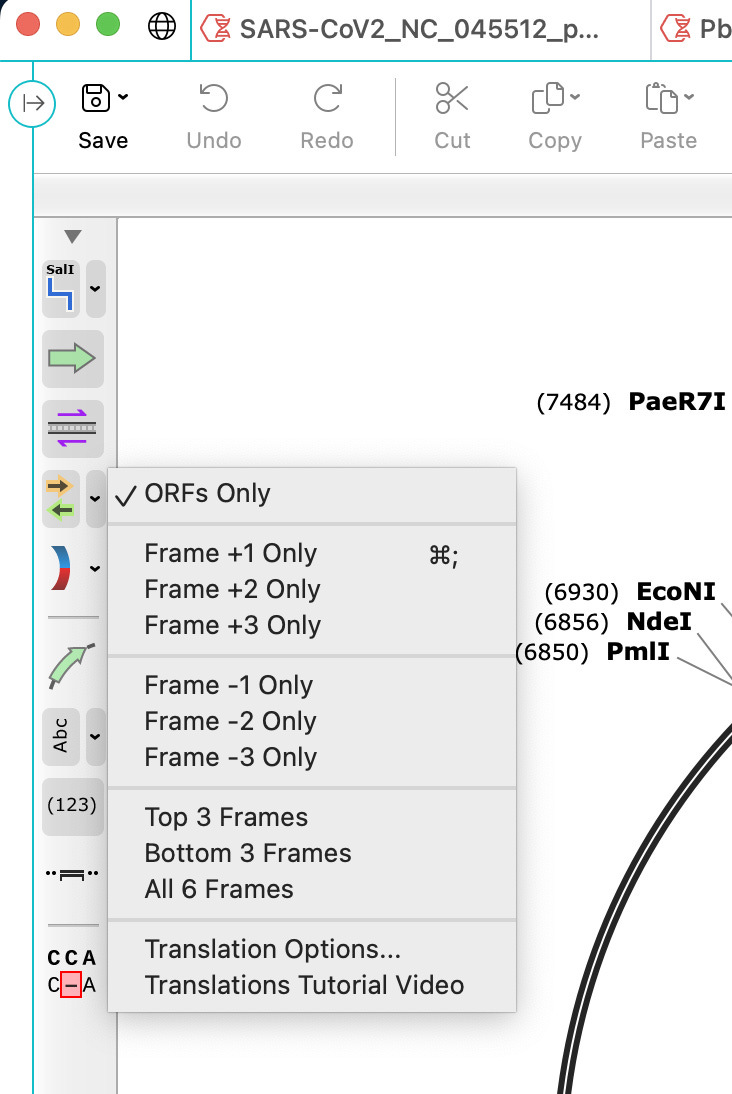
If you want to take this one step further, ask it to decorate the Open Reading Frames (ORFs) by clicking on the side panel with the gold and green arrows. Select ORFs only.
This will identify spike for you. It’s the long Golden arrow running from 6:00 to 11:30. It will also show you something that should not be there in Green running from 11:30 to 6:00. This is a mystery ORF described by Beaudoin et al. No one knows what the hell this is! NCBI BLASTP finds nothing. Its not human and if expressed will be attacked by the immune system.
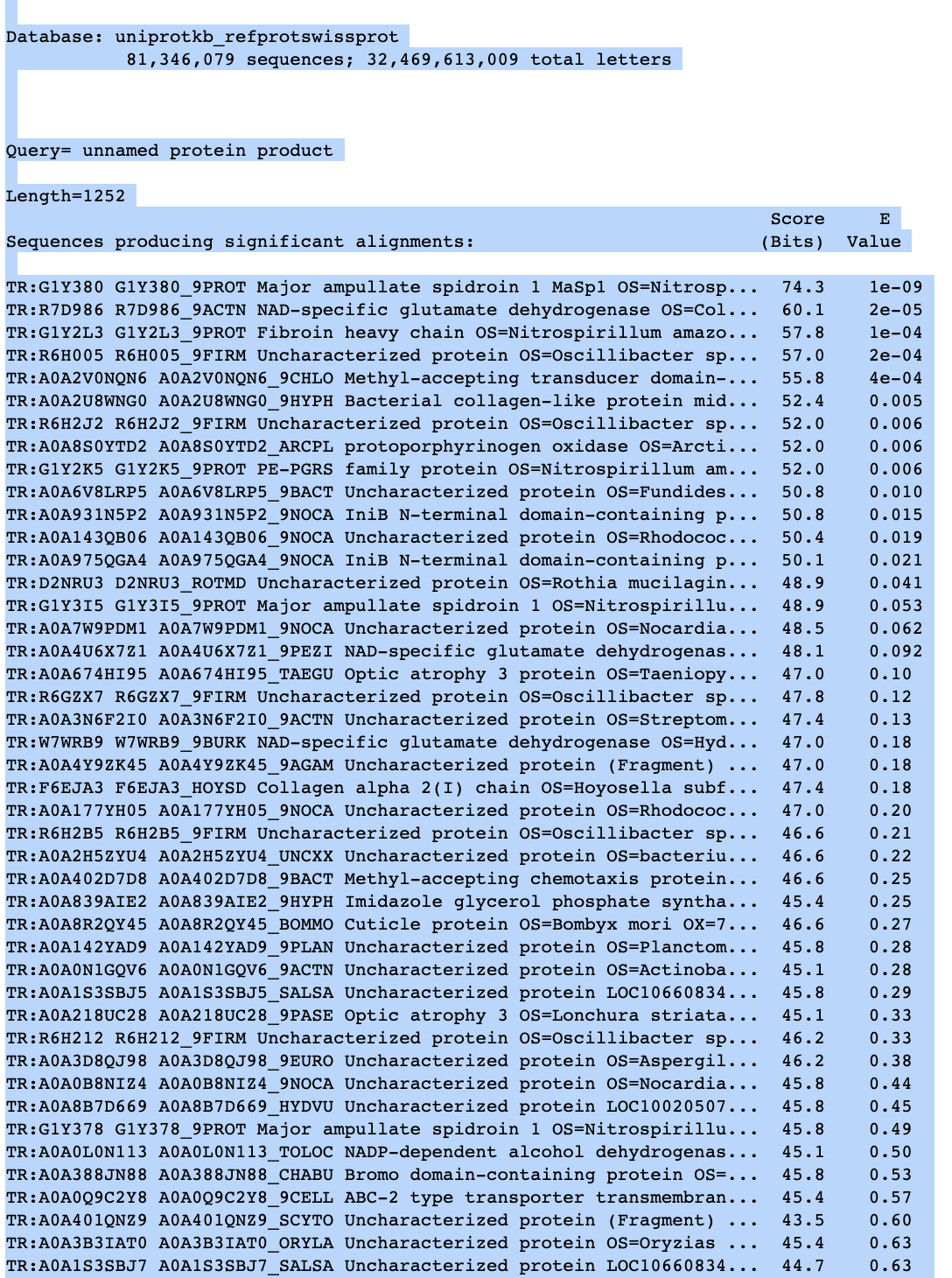
UniProt find the below hits. Not the strongest E values (E-09) but many of the hits are to proteins found in Silk, Fibroin and collagen. Hmmm.
I don’t know if this strand is expressed as RNA. The tandem 72bp SV40 Enhancer is a bidirectional promoter so odds are some RNA is made in this direction. I do see lots of Watson strand RNA (antisense sequence) in the RNA-Seq data. It should be all Crick strand RNA after the T7 promoter. I don’t know if there are Internal Ribosomal Entry Sites (IRES) as they are hard to computationally predict. I don’t see a Kozak Consensus sequence (feel free to correct me here), so maybe its a ghost but it shouldn’t be there. The viral spike doesn’t have this. This is an artifact of slipshod codon optimization or incredibly nefarious/unlucky codon optimization. It is not easy to get a 1273 ORF with a 1252 ORF on the antisense strand. This is like solving a massive sudoku puzzle during codon optimization. Maybe someone can predict the odds of this happening by chance as we dont see overlapping ORFs this long in humans?
What is this?




'someone had to actively delete this region before handing this to regulators'
game, set, and match
I actually took a break from a killer sudoku (check your spelling - no one gets this right - LOL) puzzle to read your article. I can't get away from those damned addictive puzzles and you mention it here. Argghhh. (or Arf Arf). I don't understand most of the gene stuff and microbiology, or I shy away from it for strategic reasons, but from what you wrote, there is evidence of "reckless" state of mind at the very least. Going along with the act and the resultant injury, we are in felony territory or 5 to life on the max side. I wonder if Chris has done his genealogy line.