FrameShiting Slippery Sequence
The Shift is about to hit the Fan.
By now you have probably seen a few posts that explain what Frameshifting is.
Igor Chudov has a good write up on this. By the time this publishes Jessica Rose will likely have a good write up as well.
I was also on Nick Jikomes pod cast a year ago describing our Censored PrePrint that warned about this.
This was a paper I submitted while I was quarantined with C19 ironically. Thankfully Dr. Peter McCullough of
Courageous Discourse™ with Dr. Peter McCullough & John Leake
was a co-author of the paper and offered some sound advice during this episode.
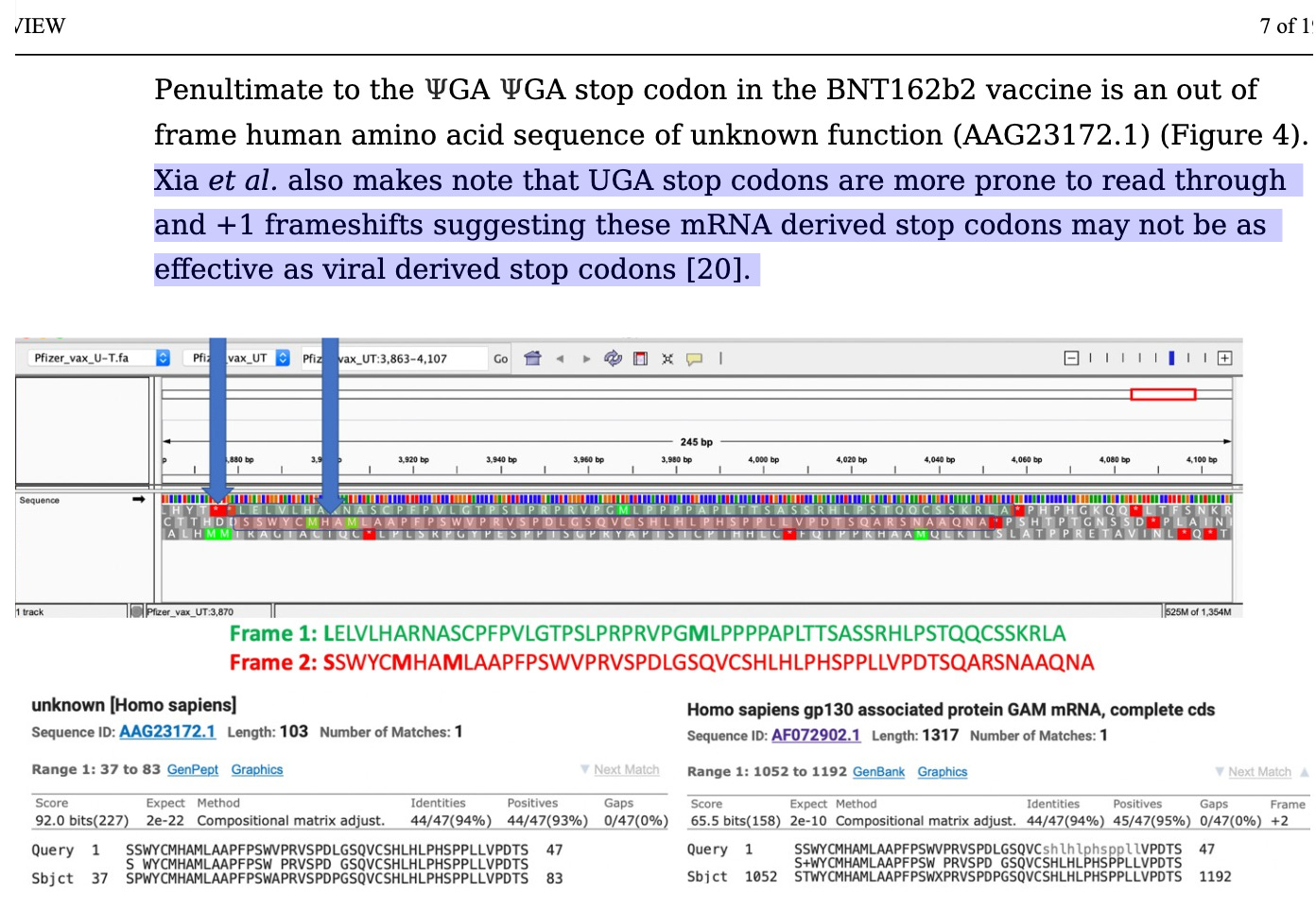
The paper spelled out this FrameShifting risk in Oct 2021. If the frameshifting occurs anywhere near the end of the sequence (3’UTR) you will skip over the stop codons and read into human peptides. Spike-Human peptides present an autoimmunity risk.
Tamara Ugolini covered this at Rebel News.
The corrupt Peer Review process was documented and published on my Substack.
Two reviewers signed off on the paper and then the Editor stepped in and buried it.
This topic was also grounds for putting me into TwitMo.
You see, the NIH was tweeting about how ‘Natural’ these modRNAs were to the public after they had filed patents on how non-natural the modRNA was. You can’t get a patent on natural RNA so the NIH had to convince the USPTO that these were NOT natural.
It turned out this was very lucrative deception as Moderna is on record paying the NIH over $400M in royalties for these non-natural modRNAs as the NIH patented one aspect of the man modification of the modRNAs.
I got TwitMo’d for pointing out the NIH was lying about these ‘Natural modRNAs’.
The Nobel Prize was also handed out to this slippery deception.
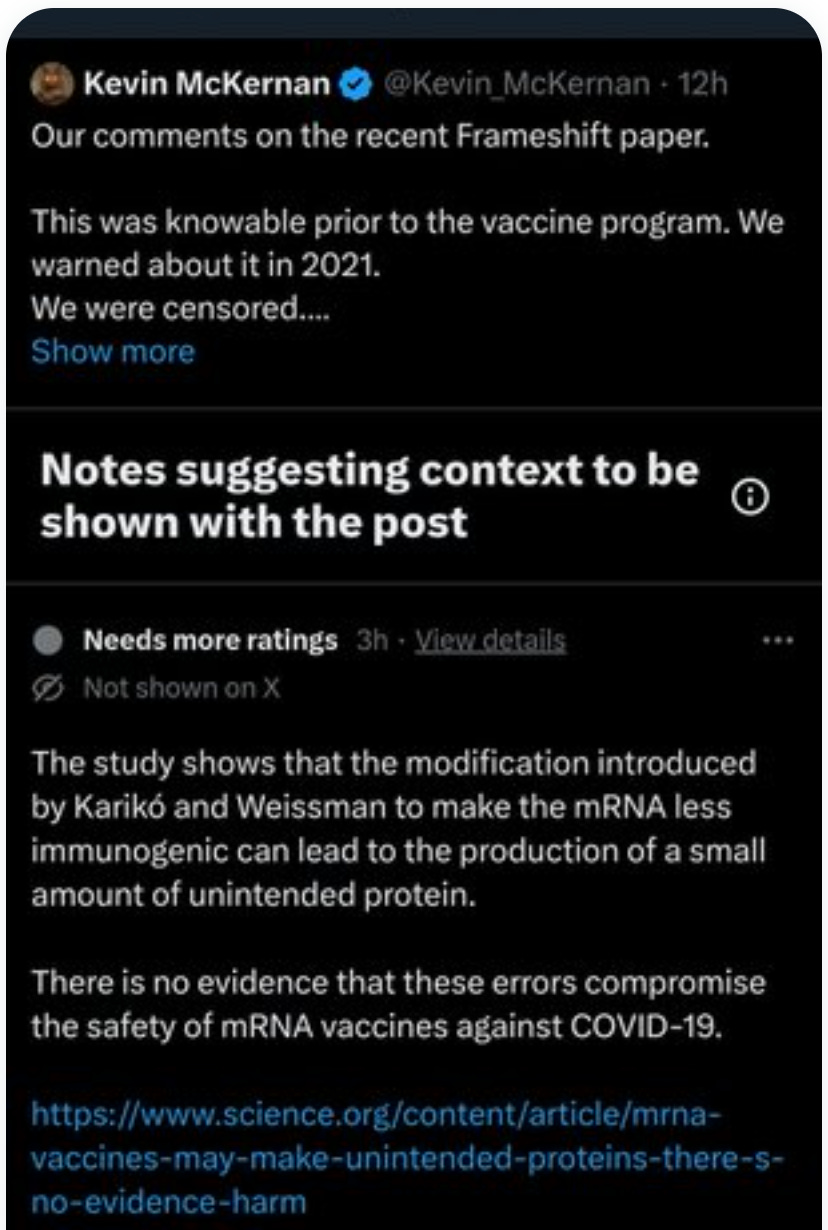
Fast forward to today, I have twitter posts about this Mulroney Frameshifting paper that are being sent to community notes by the Karen Konclave.
Today, since I was TwitMo’d over this very topic, I’m not allowed to weigh in Community Notes to defend my tweet. Fun Stuff.
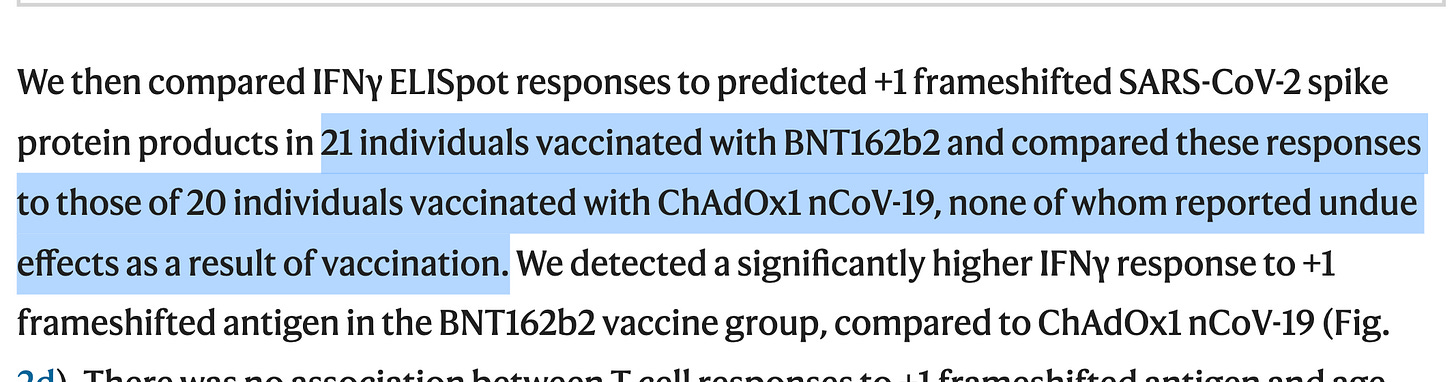
The paper being discussed this week is the shocking finding that this Frameshifting is in fact happening in 5/21 asymptomatic humans it was measured in. Repeat in the longvaxxed and I bet these numbers swell.
We submitted a comment on this Mulroney paper in Nature. Much credit to David Wiseman for so quickly putting this together and coining the phase “The shift is about to hit the fan”.
In case Nature was to censor this, we also posted it on OSF.io and ResearchGate.
Mulroney et al
It is an excellent and thorough paper but deserves several NERF awards for maintaining the safe and effective psalms despite their data being a bomb shell to the modRNA platforms.
Since others have covered this paper deeply now, I will just post a few excerpts from the paper that stood out to me below.
I don’t want to dwell on this as other have beat it to death. What I want to focus on are a weakness of this papers and more importantly 1 particular paper (Kim et al) that lied to you prior to this Mulroney study utterly destroying the prior woke stoking Kim paper.
A few notable sections below
Slippery sites are poly U tracts.
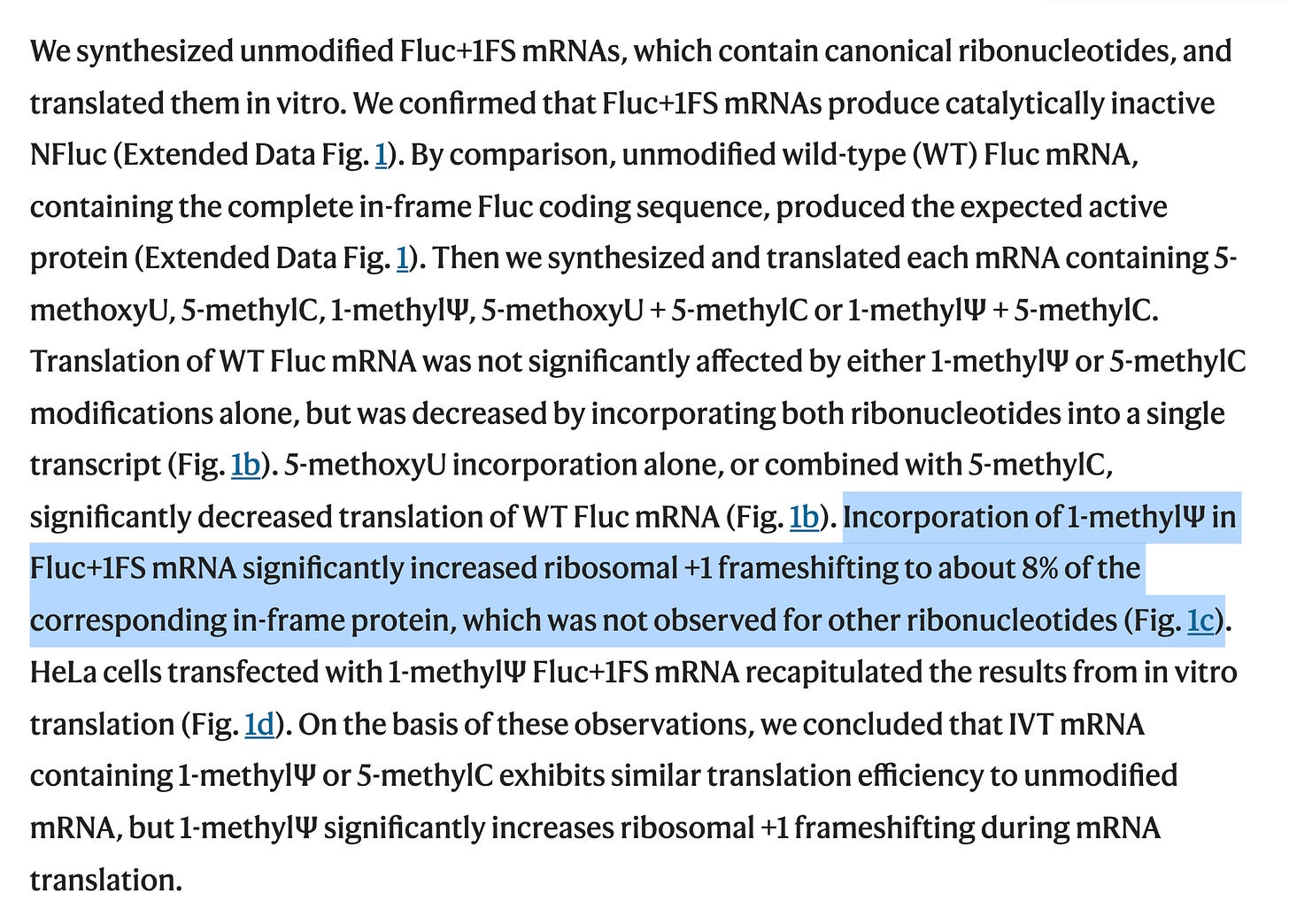
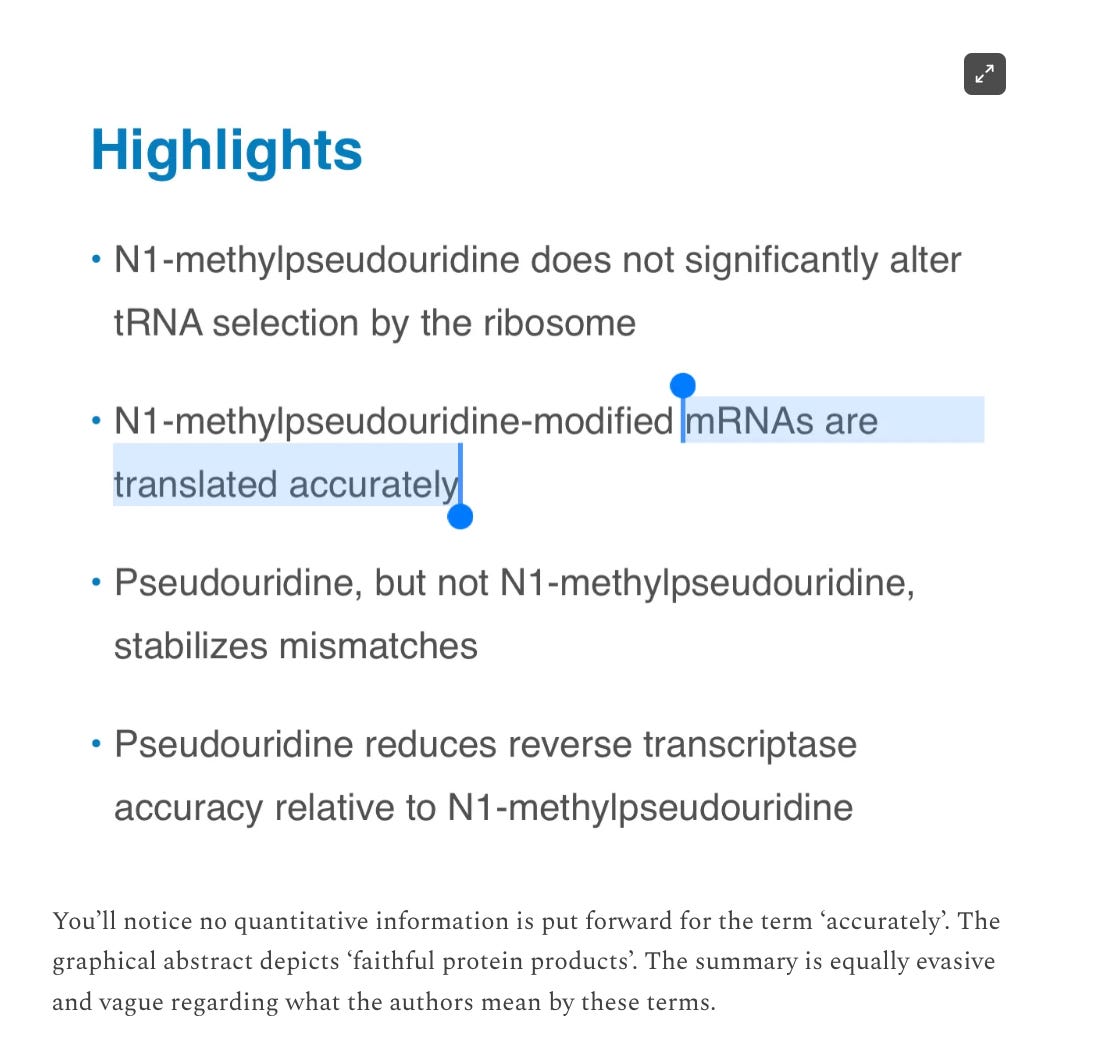
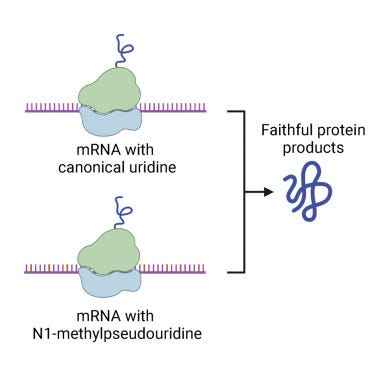
They define the slipperly sequence as tracts of 3 or more N1-methyl-pUs in a row which cause the ribosomes to stutter out of reading frame and make a mess. This is important to keep in mind as the ‘Faithful Kim paper’ was the key to the deception that filled this void for the last year. The mutton crew would regurgitate this paper like gospel.
The Faithful Kim et al paper that whitewashed this artifact in the past was torn apart in a prior substack of mine but actually put to death by Mulroney et al with hard data.
Kim was in the process of applying for a grant from NIH when he crafted this Kristian Andersen like butterball for Fauci.
What was the deception?
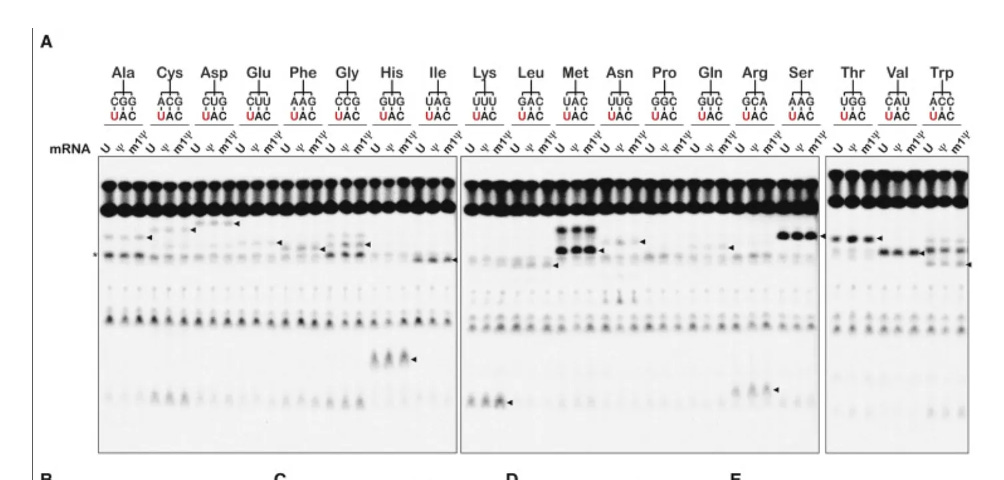
They never measured slippery sequences. They looked at single N1-methyl-pU substitutions and never challenged their assay with doublet or triplet N1-methyl-pU.
So they designed a paper that would never see it and declared it ‘Faithful’.
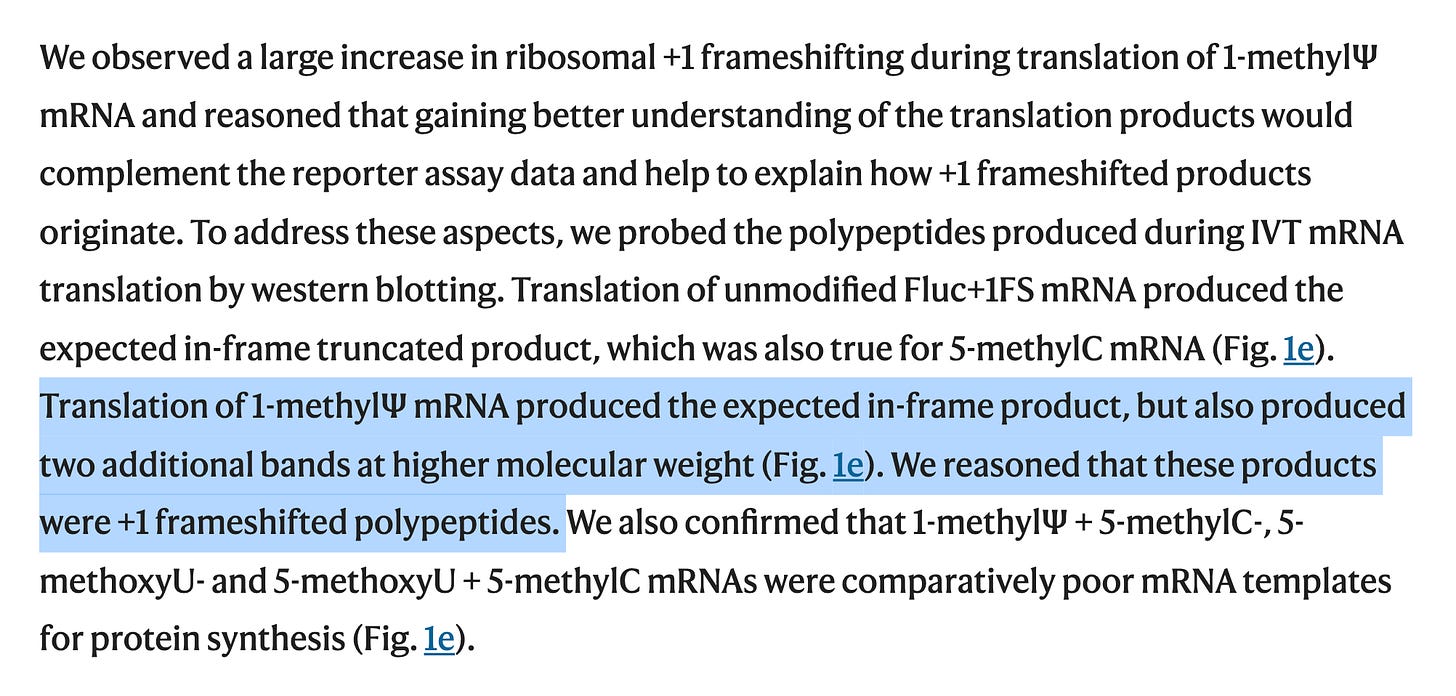
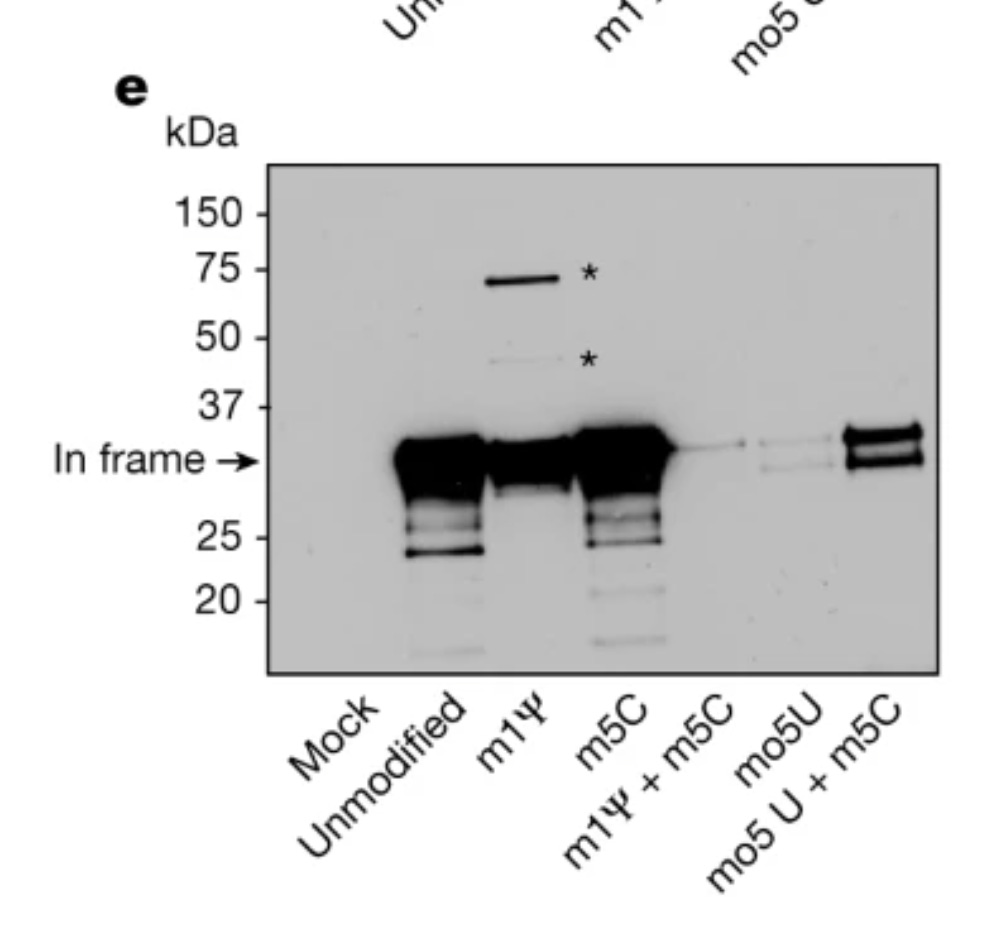
8-10% of the proteins translating are frameshifting. This is far from faithful. Its a trainwreck and likely a lower estimate of the problem.
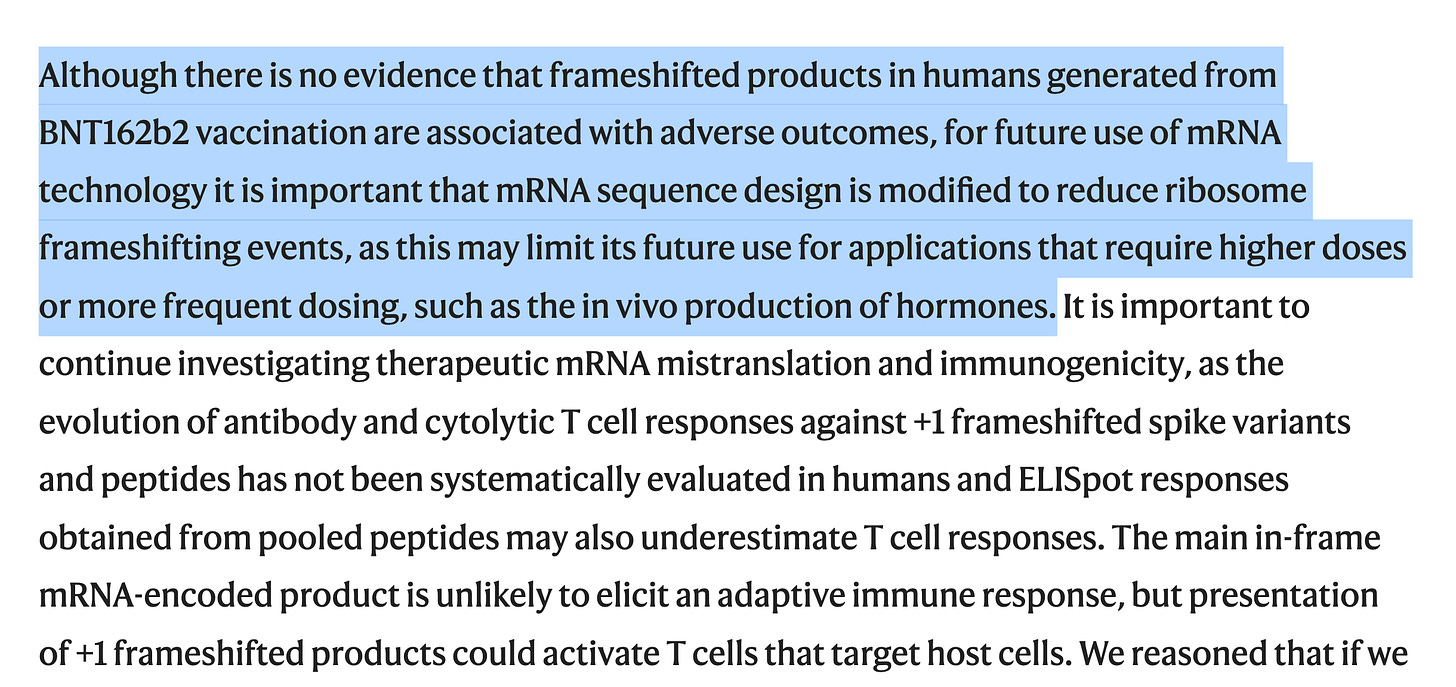
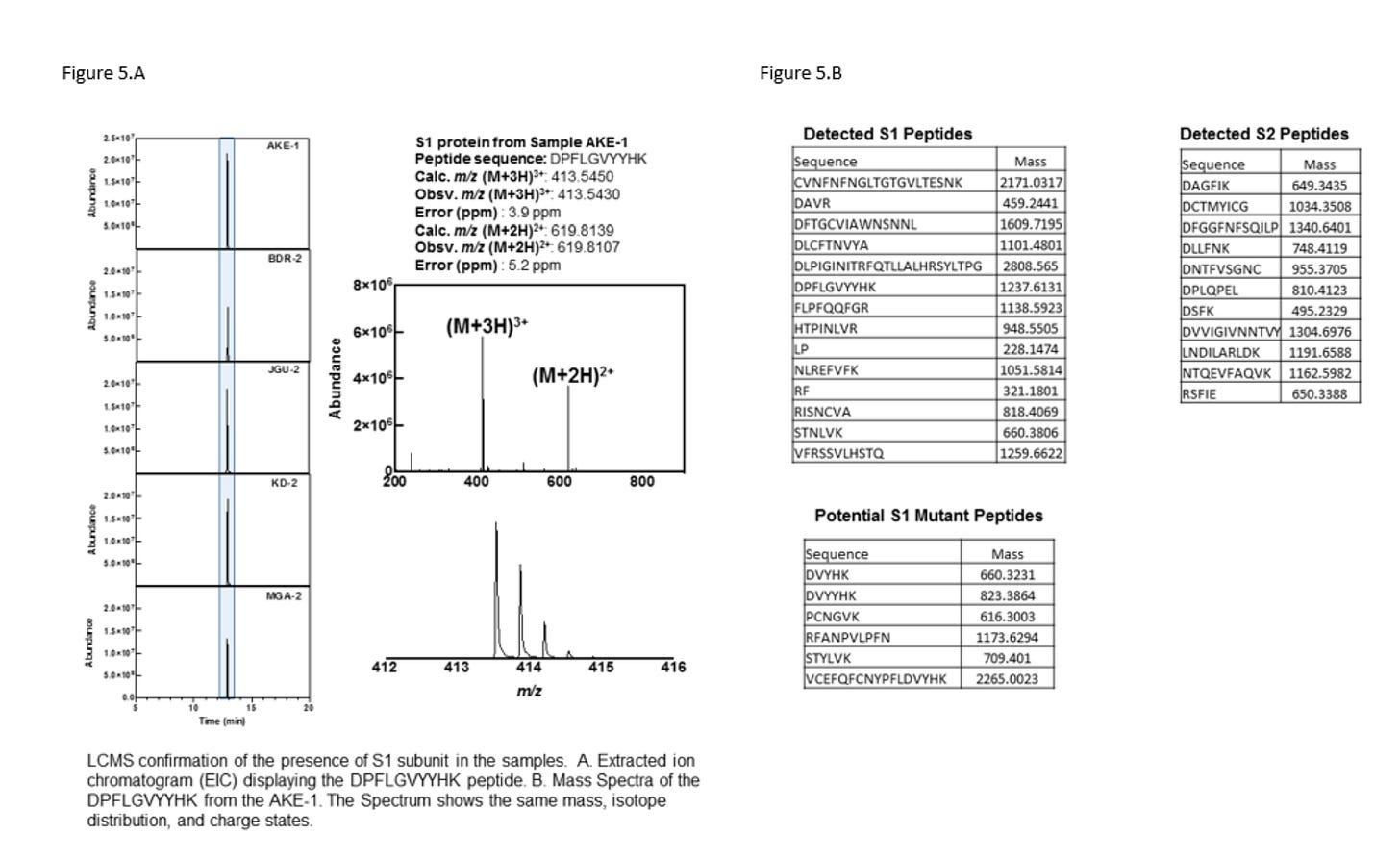
Many have claimed, this is immaterial because the immune system perfectly destroys all cells that express foreign peptides. It is so perfect at it that we have mutated S1 proteins being picked up by Bruce Pattersons lab only in the Long Vax cohort 90 days later (Figure 5B)
The immune system being so perfect that we can detect Spike proteins in exosomes 4 months later (Bansal et al) and mRNA in Breast milk 5-7 days later (Hanna et al). We also see Spike mRNA in heart tissue 30 days later described by Krauson et al.
The folks confident in this immune clearance are not paying attention to amyloidogenic peptides (nor the data that continually shows persistence of modRNA and spike protein in the long vaxxed). Amyloidogenic proteins are peptides that chain react and cause other proteins to mis-fold and become amyloidogenic. There is no safe dose. Small amounts of the prion aren’t something you count on the immune system clearing.
Its like injecting a combinatorial chemist and claiming, not to worry, the liver clears all toxic chemicals.
All this could be avoided by using non-replicative proteins as antigens. Once you start using RNA/DNA as personal vaccine factories you invite permanence into the room.
My main critique of the Mulroney paper.
They poorly addressed the transcriptional fidelity. This is step that makes the RNA from DNA. Translation is the second step.
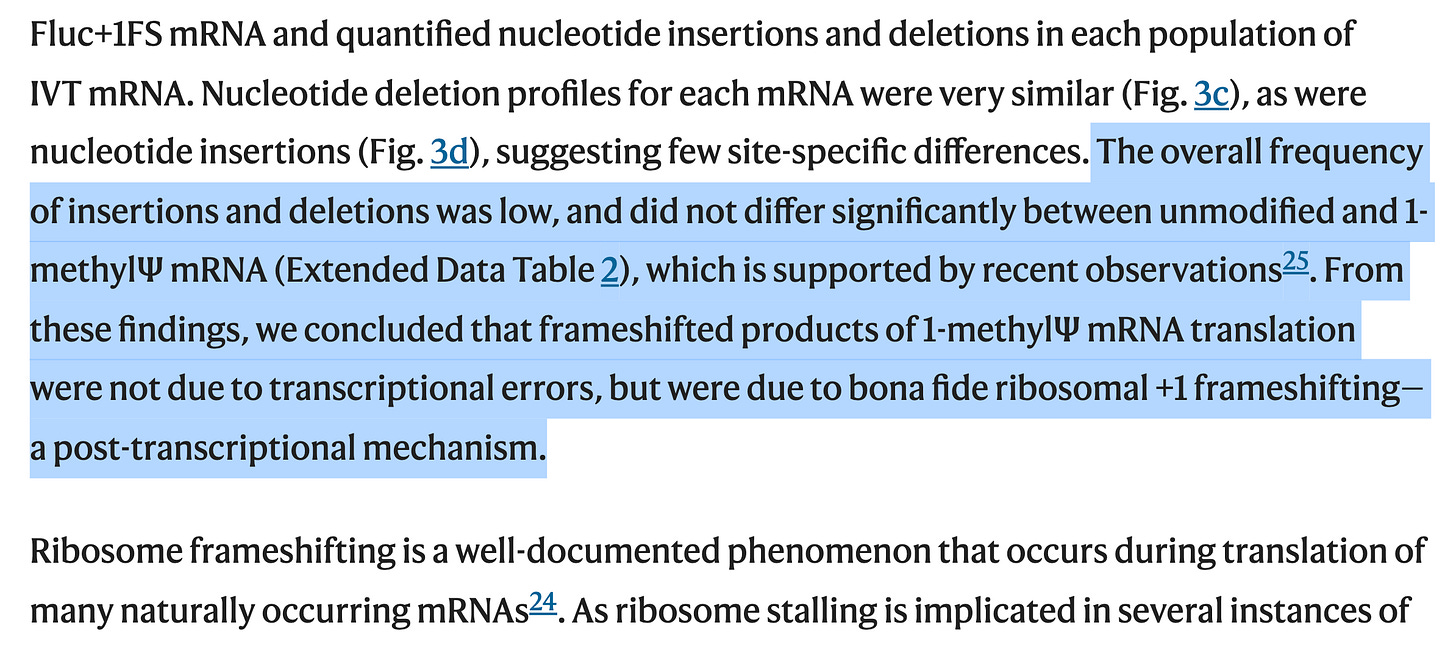
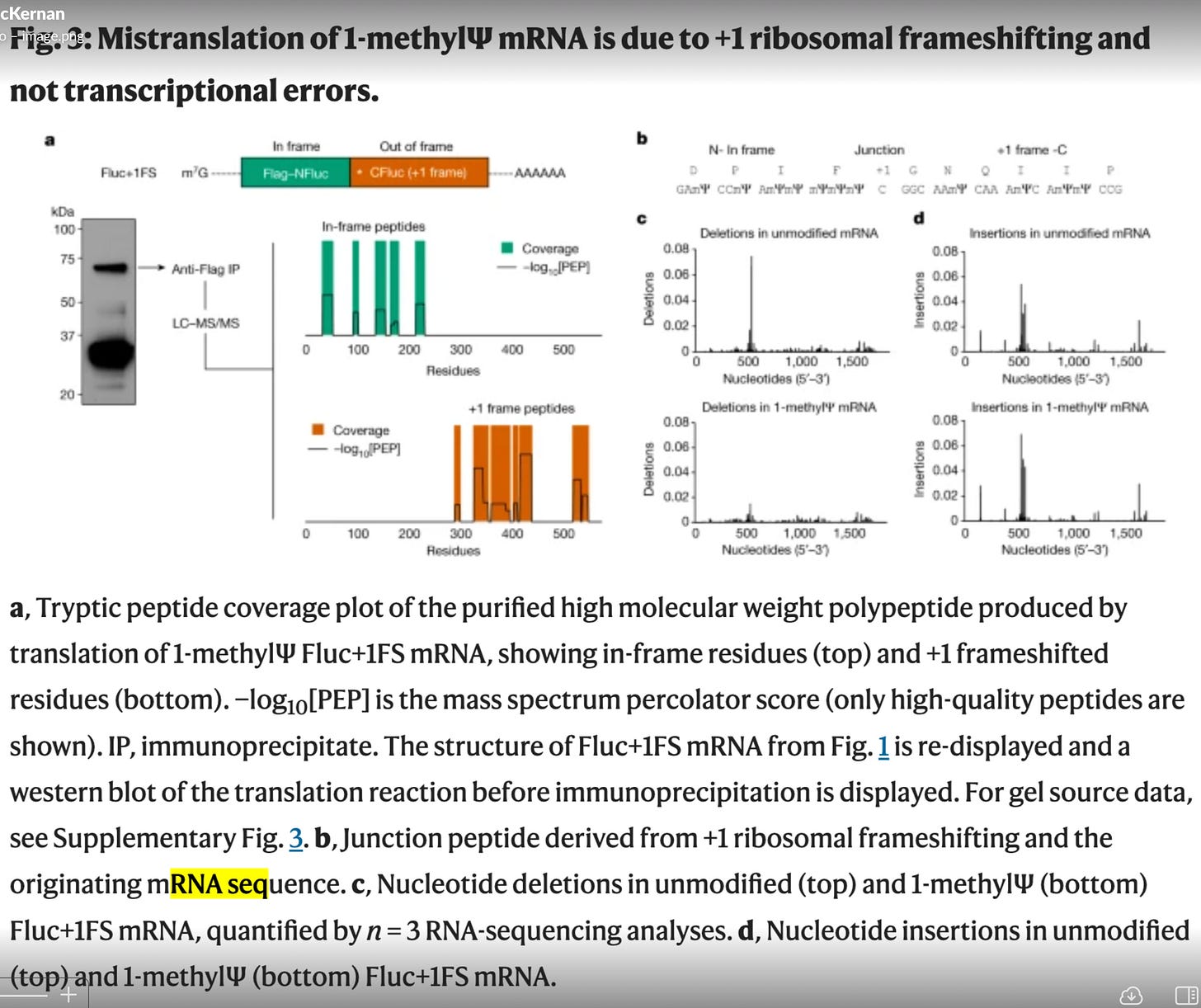
How did they do this? They used model templates from the RLuc constructs to measure this. These are only 1500-2000 bases long. The spike sequence is 4284bp long. Note figure 3c and 3d have 1,500 on the X axis. Thats the length of their RLuc construct.
The spike sequence is heavily GC enriched. Its not clear what the GC content of their model is. Its not clear what the N1-methyl-PseudoU density is in their model. If anyone finds this in the supplements, please post in the comments. I went looking and could find much. I have the RNAseq data but they only performed this on the model templates and avoided performing this sequencing on the actual vaccines? Why?
So we wil have to assemble the reads and answer these questions on our own.
Why cast such a nit pick? Because longer templates are much harder to transcribe and translate. Truncated mRNAs are commonly seen in the Moderna paper that engineered a mutated T7 polymerase to deal with this.
Likewise when your ribosomes are stalling on N1-methyl-pU. Stalling = errors. High GC and high N1-methyl-pU = more template switching which = more off target transcription and off target translation. So to make claims about transcriptional fidelity, you have to use the actual template in question. Its a pure magic trick to swap out the vaccine with a model template and declare everything is OK. In this case, its clear they had the Vaccine mRNA but chose to run this study on a mock template. That is overt deception and I would hazard to guess that they ran this experiment but didnt like what it presented and thus buried the data in favor of the mock template.
So in summary, this Frameshifting risk was known in 2021 and it was censored at the journals and on Twitter. Smoke screen papers like Kim et al appeared so some researches could get some Fauci biscuits. After billions of shots we can now see the warts through the lens of the next patent that will fix the crap forced into our kids.
And once again the goal posts begin to shift.
It stays in the arm.
Oops it doesn’t, but it doesn’t matter
It’s gone in 48hrs
Oops is still here months later, but it doesn’t matter
There is no DNA!!!
Ok there is a little, but it doesn’t matter
There is no SV40!!!
Oops there is, but it’s safe and effective
It stops transmission
Oops, it doesn’t but it makes it not so bad.
It’s good for the immunocompromised and those at risk
Oops that was never a clinical trial endpoint but its still safe and effective.














mutant proteins
.. no use trying to put lipstick on this pig
mutant proteins
Each stage of the vaccine's product development and rollout (design, testing, cooking the data, manufacturing, spreading the payola, chanting the "safe & effective" mantra) was all a huge and intentional fraud. The pharma companies knew exactly what they were doing, but believed they could get away with it. It's shocking.